Inmolecular biology,proteinsare generally thought to adopt uniquestructuresdetermined by theiramino acidsequences. However, proteins are not strictly static objects, but rather populate ensembles of (sometimes similar)conformations.Transitions between these states occur on a variety oflength scales(tenths ofangstromsto nm) and time scales (ns to s), and have been linked to functionally relevant phenomena such asallosteric signaling[1]andenzyme catalysis.[2][3]
The study ofprotein dynamicsis most directly concerned with the transitions between these states, but can also involve the nature and equilibrium populations of the states themselves. These two perspectives—kineticsandthermodynamics,respectively—can be conceptually synthesized in an "energy landscape"paradigm:[4] highly populated states and the kinetics of transitions between them can be described by the depths of energy wells and the heights of energy barriers, respectively.
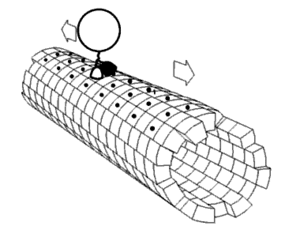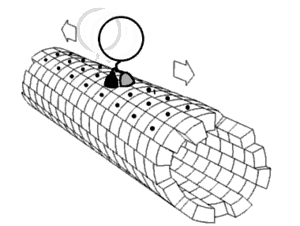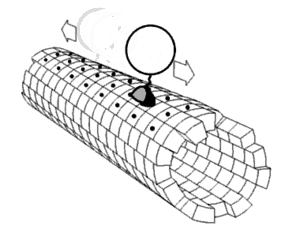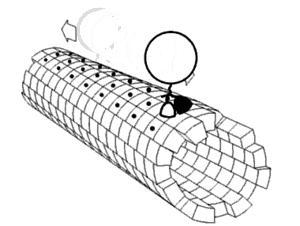
Local flexibility: atoms and residues
editPortions of protein structures often deviate from the equilibrium state. Some such excursions areharmonic,such as stochastic fluctuations ofchemical bondsand bond angles. Others areanharmonic,such as sidechains that jump between separate discrete energy minima, orrotamers.[5]
Evidence for local flexibility is often obtained fromNMR spectroscopy.Flexible and potentially disordered regions of a protein can be detected using therandom coil index.Flexibility in folded proteins can be identified by analyzing thespin relaxationof individual atoms in the protein. Flexibility can also be observed in very high-resolution electron density maps produced byX-ray crystallography,[6] particularly when diffraction data is collected at room temperature instead of the traditional cryogenic temperature (typically near 100 K).[7]Information on the frequency distribution and dynamics of local protein flexibility can be obtained usingRamanandoptical Kerr-effectspectroscopy[8]as well asanisotropic microspectroscopy[9]in the terahertz frequency domain.
Regional flexibility: intra-domain multi-residue coupling
editMany residues are in close spatial proximity in protein structures. This is true for most residues that are contiguous in the primary sequence, but also for many that are distal in sequence yet are brought into contact in the final folded structure. Because of this proximity, these residue's energy landscapes become coupled based on various biophysical phenomena such ashydrogen bonds,ionic bonds,andvan der Waals interactions(see figure).
Transitions between states for such sets of residues therefore become correlated.[10]
This is perhaps most obvious for surface-exposed loops, which often shift collectively to adopt different conformations in different crystal structures (see figure). However, coupled conformational heterogeneity is also sometimes evident in secondary structure.[11]For example, consecutive residues and residues offset by 4 in the primary sequence often interact inα helices.Also, residues offset by 2 in the primary sequence point their sidechains toward the same face ofβ sheetsand are close enough to interact sterically, as are residues on adjacent strands of the same β sheet. Some of these conformational changes are induced by post-translational modifications in protein structure, such as phosphorylation and methylation.[11][12]
When these coupled residues form pathways linking functionally important parts of a protein, they may participate inallostericsignaling. For example, when a molecule of oxygen binds to one subunit of thehemoglobintetramer, that information is allosterically propagated to the other three subunits, thereby enhancing their affinity for oxygen. In this case, the coupled flexibility in hemoglobin allows for cooperative oxygen binding, which is physiologically useful because it allows rapid oxygen loading in lung tissue and rapid oxygen unloading in oxygen-deprived tissues (e.g. muscle).
Global flexibility: multiple domains
editThe presence of multiple domains in proteins gives rise to a great deal offlexibility and mobility,leading toprotein domain dynamics.[1] Domain motions can be inferred by comparing different structures of a protein (as inDatabase of Molecular Motions), or they can be directly observed using spectra[13][2] measured byneutron spin echospectroscopy. They can also be suggested by sampling in extensive molecular dynamics trajectories[14]and principal component analysis.[15]Domain motions are important for:
- ABC transporters[16]
- adherens junction[17]
- cellular locomotion andmotor proteins[18]
- enzyme catalysis[2][19]
- formation ofprotein complexes[20]
- ion channels[21]
- mechanoreceptorsandmechanotransduction[22]
- regulatory activity[23]
- transport of metabolites acrosscell membranes[citation needed]
One of the largest observed domain motions is the 'swivelling' mechanism inpyruvate phosphate dikinase.The phosphoinositide domain swivels between two states in order to bring a phosphate group from the active site of the nucleotide binding domain to that of the phosphoenolpyruvate/pyruvate domain.[24]The phosphate group is moved over a distance of 45 Å involving a domain motion of about 100 degrees around a single residue. In enzymes, the closure of one domain onto another captures a substrate by an induced fit, allowing the reaction to take place in a controlled way. A detailed analysis by Gerstein led to the classification of two basic types of domain motion; hinge and shear.[21]Only a relatively small portion of the chain, namely the inter-domain linker and side chains undergo significant conformational changes upon domain rearrangement.[25]
Hinge motions
editA study by Hayward[26]found that the termini of α-helices and β-sheets form hinges in a large number of cases. Many hinges were found to involve two secondary structure elements acting like hinges of a door, allowing an opening and closing motion to occur. This can arise when two neighbouring strands within a β-sheet situated in one domain, diverge apart as they join the other domain. The two resulting termini then form the bending regions between the two domains. α-helices that preserve their hydrogen bonding network when bent are found to behave as mechanical hinges, storing `elastic energy' that drives the closure of domains for rapid capture of a substrate.[26]Khade et. al. worked on prediction of the hinges[27]in any conformation and further built an Elastic Network Model called hdANM[28]that can model those motions.
Helical to extended conformation
editThe interconversion of helical and extended conformations at the site of a domain boundary is not uncommon. In calmodulin, torsion angles change for five residues in the middle of a domain linking α-helix. The helix is split into two, almost perpendicular, smaller helices separated by four residues of an extended strand.[29][30]
Shear motions
editShear motions involve a small sliding movement of domain interfaces, controlled by the amino acid side chains within the interface. Proteins displaying shear motions often have a layered architecture: stacking of secondary structures. The interdomain linker has merely the role of keeping the domains in close proximity.[citation needed]
Domain motion and functional dynamics in enzymes
editThe analysis of the internal dynamics of structurally different, but functionally similar enzymes has highlighted a common relationship between the positioning of the active site and the two principal protein sub-domains. In fact, for several members of the hydrolase superfamily, the catalytic site is located close to the interface separating the two principal quasi-rigid domains.[14]Such positioning appears instrumental for maintaining the precise geometry of the active site, while allowing for an appreciable functionally oriented modulation of the flanking regions resulting from the relative motion of the two sub-domains.[citation needed]
Implications for macromolecular evolution
editEvidence suggests that protein dynamics are important for function, e.g. enzyme catalysis in dihydrofolate reductase (DHFR), yet they are also posited to facilitate the acquisition of new functions bymolecular evolution.[31] This argument suggests that proteins have evolved to have stable, mostly unique folded structures, but the unavoidable residual flexibility leads to some degree of functional promiscuity, which can be amplified/harnessed/diverted by subsequent mutations.[citation needed] Research on promiscuous proteins within theBCL-2 familyrevealed that nanosecond-scale protein dynamics can play a crucial role in protein binding behaviour and thus promiscuity.[32]
However, there is growing awareness thatintrinsically unstructured proteinsare quite prevalent in eukaryotic genomes,[33] casting further doubt on the simplest interpretation ofAnfinsen's dogma:"sequence determines structure (singular)". In effect, the new paradigm is characterized by the addition of two caveats: "sequence and cellular environment determine structural ensemble".
References
edit- ^abBu Z, Callaway DJ (2011). "Proteins move! Protein dynamics and long-range allostery in cell signaling". In Donev R (ed.).Protein Structure and Diseases.Advances in Protein Chemistry and Structural Biology. Vol. 83. Academic Press. pp.163–221.doi:10.1016/B978-0-12-381262-9.00005-7.ISBN9780123812629.PMID21570668.
- ^abcBu Z, Biehl R, Monkenbusch M, Richter D, Callaway DJ (Dec 2005)."Coupled protein domain motion in Taq polymerase revealed by neutron spin-echo spectroscopy"(PDF).Proceedings of the National Academy of Sciences of the United States of America.102(49):17646–17651.Bibcode:2005PNAS..10217646B.doi:10.1073/pnas.0503388102.PMC1345721.PMID16306270.
- ^Fraser JS, Clarkson MW, Degnan SC, Erion R, Kern D, Alber T (Dec 2009)."Hidden alternative structures of proline isomerase essential for catalysis".Nature.462(7273):669–673.Bibcode:2009Natur.462..669F.doi:10.1038/nature08615.PMC2805857.PMID19956261.
- ^ Frauenfelder H, Sligar SG, Wolynes PG (Dec 1991). "The energy landscapes and motions of proteins".Science.254(5038):1598–1603.Bibcode:1991Sci...254.1598F.doi:10.1126/science.1749933.PMID1749933.
- ^Dunbrack, Roland L (August 2002). "Rotamer Libraries in the 21st Century".Current Opinion in Structural Biology.12(4):431–440.doi:10.1016/s0959-440x(02)00344-5.PMID12163064.
- ^ Davis IW, Arendall WB, Richardson DC, Richardson JS (Feb 2006)."The backrub motion: how protein backbone shrugs when a sidechain dances".Structure.14(2):265–274.doi:10.1016/j.str.2005.10.007.PMID16472746.
- ^ Fraser JS, van den Bedem H, Samelson AJ, Lang PT, Holton JM, Echols N, Alber T (Sep 2011)."Accessing protein conformational ensembles using room-temperature X-ray crystallography".Proceedings of the National Academy of Sciences of the United States of America.108(39):16247–16252.Bibcode:2011PNAS..10816247F.doi:10.1073/pnas.1111325108.PMC3182744.PMID21918110.
- ^Turton DA, Senn HM, Harwood T, Lapthorn AJ, Ellis EM, Wynne K (June 2014)."Terahertz underdamped vibrational motion governs protein-ligand binding in solution".Nature Communications.5:3999.Bibcode:2014NatCo...5.3999T.doi:10.1038/ncomms4999.PMID24893252.
- ^Acbas, G.; Niessen, K. A.; Snell, E. H.; Markelz, A. G. (2014)."Protein Optical measurements of long-range protein vibrations".Nature Communications.5:3076.doi:10.1038/ncomms4076.PMID24430203.
- ^Bu Z, Cook J, Callaway DJ (Sep 2001). "Dynamic regimes and correlated structural dynamics in native and denatured alpha-lactalbumin".Journal of Molecular Biology.312(4):865–873.doi:10.1006/jmbi.2001.5006.PMID11575938.
- ^abCosta CH, Oliveira AR, Dos Santos AM, da Costa KS, Lima AH, Alves CN, Lameira J (October 2019). "Computational study of conformational changes in human 3-hydroxy-3-methylglutaryl coenzyme reductase induced by substrate binding".Journal of Biomolecular Structure & Dynamics.37(16):4374–4383.doi:10.1080/07391102.2018.1549508.PMID30470158.S2CID53717806.
- ^Groban ES, Narayanan A, Jacobson MP (April 2006). Shakhnovich E (ed.)."Conformational changes in protein loops and helices induced by post-translational phosphorylation".PLOS Computational Biology.2(4): e32.Bibcode:2006PLSCB...2...32G.doi:10.1371/journal.pcbi.0020032.PMC1440919.PMID16628247.
- ^Farago B, Li J, Cornilescu G, Callaway DJ, Bu Z (Nov 2010)."Activation of nanoscale allosteric protein domain motion revealed by neutron spin echo spectroscopy".Biophysical Journal.99(10):3473–3482.Bibcode:2010BpJ....99.3473F.doi:10.1016/j.bpj.2010.09.058.PMC2980739.PMID21081097.
- ^abPotestio R, Pontiggia F, Micheletti C (Jun 2009)."Coarse-grained description of protein internal dynamics: an optimal strategy for decomposing proteins in rigid subunits".Biophysical Journal.96(12):4993–5002.Bibcode:2009BpJ....96.4993P.doi:10.1016/j.bpj.2009.03.051.PMC2712024.PMID19527659.
- ^Baron R, Vellore NA (Jul 2012)."LSD1/CoREST is an allosteric nanoscale clamp regulated by H3-histone-tail molecular recognition".Proceedings of the National Academy of Sciences of the United States of America.109(31):12509–14.Bibcode:2012PNAS..10912509B.doi:10.1073/pnas.1207892109.PMC3411975.PMID22802671.
- ^Ponte-Sucre A, ed. (2009).ABC Transporters in Microorganisms.Caister Academic.ISBN978-1-904455-49-3.
- ^Callaway DJ, Nicholl ID, Shi B, Reyes G, Farago B, Bu Z (2024)."Nanoscale dynamics of the cadherin-catenin complex bound to vinculin revealed by neutron spin echo spectroscopy".Proceedings of the National Academy of Sciences of the United States of America.129(39).doi:10.1073/pnas.2408459121.PMC11441495.PMID39298480.
- ^Howard J (2001).Mechanics of motor proteins and the cytoskeleton(1st ed.). Sunderland,MA: Sinauer Associates.ISBN9780878933334.
- ^Kamerlin SC, Warshel A (May 2010)."At the dawn of the 21st century: Is dynamics the missing link for understanding enzyme catalysis?".Proteins.78(6):1339–75.doi:10.1002/prot.22654.PMC2841229.PMID20099310.
- ^Callaway DJ, Matsui T, Weiss T, Stingaciu LR, Stanley CB, Heller WT, Bu Z (Apr 2017)."Controllable Activation of Nanoscale Dynamics in a Disordered Protein Alters Binding Kinetics".Journal of Molecular Biology.429(7):987–998.doi:10.1016/j.jmb.2017.03.003.PMC5399307.PMID28285124.
- ^abGerstein M, Lesk AM, Chothia C (Jun 1994). "Structural mechanisms for domain movements in proteins".Biochemistry.33(22):6739–49.doi:10.1021/bi00188a001.PMID8204609.
- ^Nicholl ID, Matsui T, Weiss TM, Stanley CB, Heller WT, Martel A, Farago B, Callaway DJ, Bu Z (Aug 21, 2018)."Alpha-catenin structure and nanoscale dynamics in solution and in complex with F-actin".Biophysical Journal.115(4):642–654.Bibcode:2018BpJ...115..642N.doi:10.1016/j.bpj.2018.07.005.hdl:2436/621755.PMC6104293.PMID30037495.
- ^Voet D (2011).Biochemistry.Voet, Judith G. (4th ed.). Hoboken, NJ: John Wiley & Sons.ISBN9780470570951.OCLC690489261.
- ^Herzberg O, Chen CC, Kapadia G, McGuire M, Carroll LJ, Noh SJ, Dunaway-Mariano D (Apr 1996)."Swiveling-domain mechanism for enzymatic phosphotransfer between remote reaction sites".Proceedings of the National Academy of Sciences of the United States of America.93(7):2652–7.Bibcode:1996PNAS...93.2652H.doi:10.1073/pnas.93.7.2652.PMC39685.PMID8610096.
- ^Janin J, Wodak SJ (1983)."Structural domains in proteins and their role in the dynamics of protein function".Progress in Biophysics and Molecular Biology.42(1):21–78.doi:10.1016/0079-6107(83)90003-2.PMID6353481.
- ^abHayward S (Sep 1999). "Structural principles governing domain motions in proteins".Proteins.36(4):425–35.doi:10.1002/(SICI)1097-0134(19990901)36:4<425::AID-PROT6>3.0.CO;2-S.PMID10450084.S2CID29808315.
- ^Khade, Pranav M.; Kumar, Ambuj; Jernigan, Robert L. (2020-01-17)."Characterizing and Predicting Protein Hinges for Mechanistic Insight".Journal of Molecular Biology.432(2):508–522.doi:10.1016/j.jmb.2019.11.018.ISSN1089-8638.PMC7029793.PMID31786268.
- ^Khade, Pranav M.; Scaramozzino, Domenico; Kumar, Ambuj; Lacidogna, Giuseppe; Carpinteri, Alberto; Jernigan, Robert L. (2021-11-16)."hdANM: a new comprehensive dynamics model for protein hinges".Biophysical Journal.120(22):4955–4965.doi:10.1016/j.bpj.2021.10.017.ISSN1542-0086.PMC8633836.PMID34687719.
- ^Meador WE, Means AR, Quiocho FA (Aug 1992). "Target enzyme recognition by calmodulin: 2.4 A structure of a calmodulin-peptide complex".Science.257(5074):1251–1255.Bibcode:1992Sci...257.1251M.doi:10.1126/science.1519061.PMID1519061.
- ^Ikura M, Clore GM, Gronenborn AM, Zhu G, Klee CB, Bax A (May 1992). "Solution structure of a calmodulin-target peptide complex by multidimensional NMR".Science.256(5057):632–638.Bibcode:1992Sci...256..632I.doi:10.1126/science.1585175.PMID1585175.
- ^ Tokuriki N, Tawfik DS (Apr 2009). "Protein dynamism and evolvability".Science.324(5924):203–207.Bibcode:2009Sci...324..203T.doi:10.1126/science.1169375.PMID19359577.S2CID28576496.
- ^Heckmeier, Philipp J.; Ruf, Jeannette; Janković, Brankica G.; Hamm, Peter (7 March 2023). "MCL-1 promiscuity and the structural resilience of its binding partners".The Journal of Chemical Physics.158(9).arXiv:2211.08934.doi:10.1063/5.0137239.
- ^ Dyson HJ,Wright PE (Mar 2005). "Intrinsically unstructured proteins and their functions".Nature Reviews Molecular Cell Biology.6(3):197–208.doi:10.1038/nrm1589.PMID15738986.S2CID18068406.