α-Amylaseis anenzyme(EC3.2.1.1;systematic name4-α-D-glucan glucanohydrolase) thathydrolysesα bonds of large, α-linkedpolysaccharides,such asstarchandglycogen,yielding shorter chains thereof,dextrins,andmaltose,through the following biochemical process:[2]
| GH13 catalytic domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Cyclodextrin glucanotransferase (e.c.2.4.1.19) (cgtase) | |||||||||
| Identifiers | |||||||||
| Symbol | Alpha-amylase | ||||||||
| Pfam | PF00128 | ||||||||
| Pfamclan | CL0058 | ||||||||
| InterPro | IPR006047 | ||||||||
| SCOP2 | 1ppi/SCOPe/SUPFAM | ||||||||
| OPM superfamily | 117 | ||||||||
| OPM protein | 1wza | ||||||||
| CAZy | GH13 | ||||||||
| CDD | cd11338 | ||||||||
| |||||||||
| Alpha-amylase C-terminal beta-sheet domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
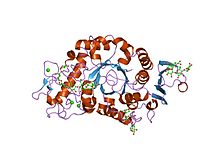 Crystal structure of barley Alpha -amylase isozyme 1 (amy1) inactive mutant d180a in complex with maltoheptaose | |||||||||
| Identifiers | |||||||||
| Symbol | Alpha-amyl_C2 | ||||||||
| Pfam | PF07821 | ||||||||
| InterPro | IPR012850 | ||||||||
| |||||||||
| Alpha amylase, C-terminal all-beta domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
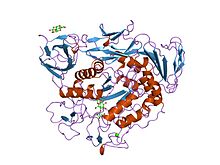 maltotriose complex of preconditioned cyclodextrin glycosyltransferase mutant | |||||||||
| Identifiers | |||||||||
| Symbol | Alpha-amylase_C | ||||||||
| Pfam | PF02806 | ||||||||
| Pfamclan | CL0369 | ||||||||
| InterPro | IPR006048 | ||||||||
| SCOP2 | 1ppi/SCOPe/SUPFAM | ||||||||
| |||||||||
- Endohydrolysis of (1→4)-α-D-glucosidic linkages in polysaccharides containing three or more (1→4)-α-linkedD-glucose units
It is the major form ofamylasefound in humans and other mammals.[3]It is also present in seeds containing starch as a food reserve, and is secreted by many fungi. It is a member ofglycoside hydrolase family 13.
In human biology
editAlthough found in many tissues,amylaseis most prominent inpancreatic juiceandsaliva,each of which has its ownisoformof human α-amylase. They behave differently onisoelectric focusing,and can also be separated in testing by using specificmonoclonal antibodies.In humans, all amylase isoforms link tochromosome 1p21 (seeAMY1A).
Salivary amylase (ptyalin)
editAmylase is found in saliva and breaksstarchintomaltoseanddextrin.This form of amylase is also called "ptyalin"/ˈtaɪəlɪn/,which was named by chemistJöns Jacob Berzelius.The name derives from the Greek word πτυω (I spit), because the substance was obtained from saliva.[4]It will break large, insoluble starch molecules into soluble starches (amylodextrin,erythrodextrin,andachrodextrin) producing successively smaller starches and ultimatelymaltose.Ptyalin acts on linear α(1,4)glycosidic linkages,but compoundhydrolysisrequires an enzyme that acts on branched products. Salivary amylase is inactivated in thestomachbygastric acid.In gastric juice adjusted to pH 3.3, ptyalin was totally inactivated in 20 minutes at 37 °C. In contrast, 50% of amylase activity remained after 150 minutes of exposure to gastric juice at pH 4.3.[5]Both starch, the substrate for ptyalin, and the product (short chains of glucose) are able to partially protect it against inactivation by gastric acid. Ptyalin added to buffer at pH 3.0 underwent complete inactivation in 120 minutes; however, addition of starch at a 0.1% level resulted in 10% of the activity remaining, and similar addition of starch to a 1.0% level resulted in about 40% of the activity remaining at 120 minutes.[6]
Optimum conditions for ptyalin
edit- OptimumpH– 7.0;[7]5.6-6.9[8]
- Human body temperature- 37 degrees Celsius[8]
- Presence of certainanionsand activators:
Genetic variation in human salivary amylase
editThe salivary amylase gene has undergone duplication during evolution, and DNA hybridization studies indicate many individuals have multiple tandem repeats of the gene. The number of gene copies correlates with the levels of salivary amylase, as measured by protein blot assays using antibodies to human amylase. Gene copy number is associated with apparent evolutionary exposure to high-starch diets.[9]For example, a Japanese individual had 14 copies of the amylase gene (one allele with 10 copies, and a second allele with four copies). The Japanese diet has traditionally contained large amounts ofricestarch. In contrast, a Biaka individual carried six copies (three copies on each allele). TheBiakaare rainforest hunter-gatherers who have traditionally consumed a low-starch diet. Perry and colleagues speculated the increased copy number of the salivary amylase gene may have enhanced survival coincident to a shift to a starchy diet during human evolution.
Pancreatic amylase
editPancreatic α-amylase randomly cleaves theα(1-4) glycosidic linkagesofamyloseto yielddextrin,maltose,ormaltotriose.It adopts a double displacement mechanism with retention ofanomeric configuration.In humans, the salivary amylase evolved from a copy of it.[9]
In pathology
editThe test for amylase is easier to perform than that forlipase,making it the primary test used to detect and monitorpancreatitis.Medical laboratorieswill usually measure either pancreatic amylase or total amylase. If only pancreatic amylase is measured, an increase will not be noted withmumpsor other salivary gland trauma.
However, because of the small amount present, timing is critical when samplingbloodfor this measurement. Blood should be taken soon after a bout of pancreatitis pain, otherwise it isexcretedrapidly by thekidneys.
Salivary α-amylase has been used as abiomarkerforstress[10][11]and as a surrogate marker ofsympathetic nervous system (SNS)activity[12]that does not require a blood draw.
Interpretation
editIncreased plasma levels in humans are found in:
- Salivarytrauma (includinganaestheticintubation)
- Mumps– due toinflammationof thesalivary glands
- Pancreatitis– because of damage to the cells that produce amylase
- Kidney failure– due to reduced excretion
Total amylase readings of over 10 times the upper limit of normal (ULN) are suggestive of pancreatitis. Five to 10 times the ULN may indicateileusorduodenaldisease or kidney failure, and lower elevations are commonly found in salivary gland disease.
Genes
editIn grain
editα-Amylase activity in grain is measured by, for instance, theHagberg–Perten Falling Number,a test to assess sprout damages,[13]or thePhadebasmethod. It occurs inwheat.[14]
Industrial use
editα-Amylase is used in ethanol production to break starches in grains into fermentable sugars.
The first step in the production ofhigh-fructose corn syrupis the treatment ofcornstarchwith α-amylase, which cleaves the long starch polymers into shorter chains ofoligosaccharides.
An α-amylase called "Termamyl", sourced fromBacillus licheniformis,is also used in some detergents, especially dishwashing and starch-removing detergents.[15]
Seeamylasefor more uses of the amylase family in general.
Potential for medical use
editα-Amylase has exhibited efficacy in degrading polymicrobial bacterialbiofilmsby hydrolyzing theα(1→4) glycosidic linkageswithin the structural, matrix exopolysaccharides of theextracellular polymeric substance(EPS).[16][17]
Disease and health relevance
edit- Diabetes:α-glucosidase and Alpha -amylase inhibitors are found in several raw plants/herbs such ascinnamon[18]and bacteria containingacarbose[19]They are used as anti-diabetic drugs. The intake of a single dose of before a meal containing complex carbohydrates clearly suppresses the glucose spike and may decrease the postprandial hyperglycemia (higher than 140 mg/dL; >7.8 mmol/L) in patients with type II diabetes.[18]
Buffer inhibition
editThetrismolecule is reported to inhibit a number of bacterial α-amylases,[20][21]so they should not be used in tris buffer.
Determination
editSeveral methods are available for determination of α-amylase activity, and different industries tend to rely on different methods. The starch iodine test, a development of theiodine test,is based on colour change, as α-amylase degrades starch and is commonly used in many applications. A similar but industrially produced test is thePhadebasamylase test, which is used as a qualitative and quantitative test within many industries, such as detergents, various flour, grain, and malt foods, and forensic biology.
Modifiedcolorimetricmicrodetermination of amylase is described in which the digestion of starch is measured by the decrease in the starch-iodine color.[22]
Domain architecture
editα-Amylases contain a number of distinct protein domains. Thecatalyticdomain has astructureconsisting of an eight-stranded α/β barrel that contains the active site, interrupted by a ~70-amino acidcalcium-binding domain protruding betweenβ-strand3 andα-helix3, and a carboxyl-terminal Greek keyβ-barreldomain.[23]Several α-amylases contain a β-sheet domain, usually at the C terminus. This domain is organised as a five-stranded antiparallel β-sheet.[24][25]Several α-amylases contain an all-β domain, usually at the C terminus.[26]
See also
editReferences
edit- ^Ramasubbu N, Paloth V, Luo Y, Brayer GD, Levine MJ (May 1996)."Structure of human salivary α-amylase at 1.6 Å resolution: implications for its role in the oral cavity".Acta Crystallographica D.52(Pt 3): 435–46.doi:10.1107/S0907444995014119.PMID15299664.
- ^Kierulf P."Amylase".Store Medisinske Leksikon.Store Norske Leksikon.Retrieved24 January2021.
- ^Voet D, Voet JG (2005).Biochimie(2nd ed.). Bruxelles: De Boeck. p. 1583.
- ^J. Berzelius (Ms. Esslinger, trans.),Traité de Chimie(Paris, France: Firmin Didot Frerès, 1833), vol. 7,page 156.
- ^Fried M, Abramson S, Meyer JH (October 1987). "Passage of salivary amylase through the stomach in humans".Digestive Diseases and Sciences.32(10): 1097–103.doi:10.1007/bf01300195.PMID3652896.S2CID24845837.
- ^Rosenblum JL, Irwin CL, Alpers DH (May 1988). "Starch and glucose oligosaccharides protect salivary-type amylase activity at acid pH".The American Journal of Physiology.254(5 Pt 1): G775–80.doi:10.1152/ajpgi.1988.254.5.G775.PMID2452576.
- ^"Amylase, Alpha – Worthington Enzyme Manual".worthington-biochem.Archivedfrom the original on 14 October 2016.
- ^abValls, Cristina; Rojas, Cristina; Pujadas, Gerard; Garcia-Vallve, Santi; Mulero, Miquel (July 2012)."Characterization of the activity and stability of amylase from saliva and detergent: Laboratory practicals for studying the activity and stability of amylase from saliva and various commercial detergents".Biochemistry and Molecular Biology Education.40(4): 254–265.doi:10.1002/bmb.20612.PMID22807429.S2CID36680999.
- ^abPerry GH, Dominy NJ, Claw KG, Lee AS, Fiegler H, Redon R, Werner J, Villanea FA, Mountain JL, Misra R, Carter NP, Lee C, Stone AC (October 2007)."Diet and the evolution of human amylase gene copy number variation".Nature Genetics.39(10): 1256–60.doi:10.1038/ng2123.PMC2377015.PMID17828263.
- ^Noto Y, Sato T, Kudo M, Kurata K, Hirota K (December 2005)."The relationship between salivary biomarkers and state-trait anxiety inventory score under mental arithmetic stress: a pilot study".Anesthesia and Analgesia.101(6): 1873–6.doi:10.1213/01.ANE.0000184196.60838.8D.PMID16301277.S2CID22252878.
- ^Granger DA, Kivlighan KT, el-Sheikh M, Gordis EB, Stroud LR (March 2007). "Salivary α-amylase in biobehavioral research: recent developments and applications".Annals of the New York Academy of Sciences.1098(1): 122–44.Bibcode:2007NYASA1098..122G.doi:10.1196/annals.1384.008.PMID17332070.S2CID222075003.
- ^Nater UM, Rohleder N (May 2009). "Salivary α-amylase as a non-invasive biomarker for the sympathetic nervous system: current state of research".Psychoneuroendocrinology.34(4): 486–96.doi:10.1016/j.psyneuen.2009.01.014.PMID19249160.S2CID7564969.
- ^ "Falling Number – Introduction".Perten Instruments. 2005. Archived fromthe originalon 9 September 2009.Retrieved21 November2009.
- ^Gatehouse AM, Davison GM, Newell CA, Merryweather A, Hamilton WD, Burgess EP, Gilbert RJ, Gatehouse JA (1997). "Transgenic potato plants with enhanced resistance to the tomato moth,Lacanobia oleracea:growth room trials ".Molecular Breeding.3(1).Springer Science+Business:49–63.doi:10.1023/a:1009600321838.ISSN1380-3743.S2CID23765916.
- ^ "The use of enzymes in detergents".Faculty of Engineering, Science and the Built Environment, London South Bank University. 20 December 2004. Archived fromthe originalon 20 October 2009.Retrieved21 November2009.
- ^Fleming D, Rumbaugh KP (April 2017)."Approaches to Dispersing Medical Biofilms".Microorganisms.5(2): 15.doi:10.3390/microorganisms5020015.PMC5488086.PMID28368320.
- ^Fleming D, Chahin L, Rumbaugh K (February 2017)."Glycoside Hydrolases Degrade Polymicrobial Bacterial Biofilms in Wounds".Antimicrobial Agents and Chemotherapy.61(2): AAC.01998–16.doi:10.1128/AAC.01998-16.PMC5278739.PMID27872074.
- ^abMoreira, Fernanda Duarte; Reis, Caio Eduardo Gonçalves; Gallassi, Andrea Donatti; Moreira, Daniel Carneiro; Welker, Alexis Fonseca (9 October 2024). Dardari, Dured (ed.)."Suppression of the postprandial hyperglycemia in patients with type 2 diabetes by a raw medicinal herb powder is weakened when consumed in ordinary hard gelatin capsules: A randomized crossover clinical trial".PLOS ONE.19(10): e0311501.doi:10.1371/journal.pone.0311501.ISSN1932-6203.PMC11463819.PMID39383145.
- ^Hayward, Nicholas J.; McDougall, Gordon J.; Farag, Sara; Allwood, J. William; Austin, Ceri; Campbell, Fiona; Horgan, Graham; Ranawana, Viren (December 2019)."Cinnamon Shows Antidiabetic Properties that Are Species-Specific: Effects on Enzyme Activity Inhibition and Starch Digestion".Plant Foods for Human Nutrition.74(4): 544–552.doi:10.1007/s11130-019-00760-8.ISSN0921-9668.PMC6900266.PMID31372918.
- ^Ghalanbor Z, Ghaemi N, Marashi SA, Amanlou M, Habibi-Rezaei M, Khajeh K, Ranjbar B (2008). "Binding of Tris to Bacillus licheniformis Alpha -amylase can affect its starch hydrolysis activity".Protein and Peptide Letters.15(2): 212–4.doi:10.2174/092986608783489616.PMID18289113.
- ^Aghajari N, Feller G, Gerday C, Haser R (March 1998)."Crystal structures of the psychrophilic α-amylase from Alteromonas haloplanctis in its native form and complexed with an inhibitor".Protein Science.7(3): 564–72.doi:10.1002/pro.5560070304.PMC2143949.PMID9541387.
- ^Pimstone, Neville R. (1964)."A Study of the Starch-Iodine Complex: A Modified Colourimetric Micro Determination of Amylase in Biological Fluids".Clinical Chemistry.10(10).American Association for Clinical Chemistry:891–906.doi:10.1093/clinchem/10.10.891.Archivedfrom the original on 14 May 2022.
- ^Abe A, Yoshida H, Tonozuka T, Sakano Y, Kamitori S (December 2005)."Complexes ofThermoactinomyces vulgarisR-47 Alpha -amylase 1 and pullulan model oligossacharides provide new insight into the mechanism for recognizing substrates with α-(1,6) glycosidic linkages ".The FEBS Journal.272(23): 6145–53.doi:10.1111/j.1742-4658.2005.05013.x.PMID16302977.S2CID41008169.
- ^Kadziola A, Søgaard M, Svensson B, Haser R (April 1998). "Molecular structure of a barley Alpha -amylase-inhibitor complex: implications for starch binding and catalysis".Journal of Molecular Biology.278(1): 205–17.doi:10.1006/jmbi.1998.1683.PMID9571044.
- ^Kadziola A, Abe J, Svensson B, Haser R (May 1994). "Crystal and molecular structure of barley α-amylase".Journal of Molecular Biology.239(1): 104–21.doi:10.1006/jmbi.1994.1354.PMID8196040.
- ^Machius M, Wiegand G, Huber R (March 1995). "Crystal structure of calcium-depletedBacillus licheniformisα-amylase at 2.2 Å resolution ".Journal of Molecular Biology.246(4): 545–59.doi:10.1006/jmbi.1994.0106.PMID7877175.
External links
edit- The Alpha -Amylase Protein
- Alpha -Amylaseat the U.S. National Library of MedicineMedical Subject Headings(MeSH)
