Cytochrome c oxidase
| Cytochrome c oxidase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC no. | 1.9.3.1 | ||||||||
| CAS no. | 9001-16-5 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDBstructures | RCSB PDBPDBePDBsum | ||||||||
| Gene Ontology | AmiGO/QuickGO | ||||||||
| |||||||||
| Cytochrome c oxidase | |
|---|---|
 | |
| Identifiers | |
| Symbol | Cytochrome c oxidase |
| OPM superfamily | 4 |
| OPM protein | 2dyr |
| Membranome | 257 |
Theenzymecytochrome c oxidaseorComplex IV(wasEC1.9.3.1,now reclassified as a translocaseEC 7.1.1.9) is a largetransmembrane proteincomplex found inbacteria,archaea,and themitochondriaofeukaryotes.[1]
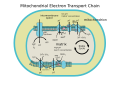
It is the last enzyme in therespiratoryelectron transport chainofcellslocated in themembrane.It receives an electron from each of fourcytochrome cmolecules and transfers them to oneoxygenmolecule and fourprotons,producing two molecules of water. In addition to binding the four protons from the inner aqueous phase, it transports another four protons across the membrane, increasing the transmembrane difference of protonelectrochemical potential,which theATP synthasethen uses to synthesizeATP.
Structure
[edit]The complex
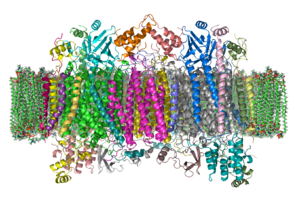
[edit]The complex is a largeintegral membrane proteincomposed of severalmetal prosthetic sitesand 14[2]protein subunits in mammals. In mammals, eleven subunits are nuclear in origin, and three are synthesized in the mitochondria. The complex contains twohemes,acytochrome aandcytochrome a3,and two copper centers, the CuAand CuBcenters.[3]In fact, the cytochrome a3and CuBform a binuclear center that is the site of oxygen reduction.Cytochrome c,which is reduced by the preceding component of the respiratory chain (cytochrome bc1 complex, Complex III), docks near the CuAbinuclear center and passes an electron to it, being oxidized back to cytochrome c containing Fe3+.The reduced CuAbinuclear center now passes an electron on to cytochrome a, which in turn passes an electron on to the cytochrome a3>-CuBbinuclear center. The two metal ions in this binuclear center are 4.5 Å apart and coordinate ahydroxide ionin the fully oxidized state.
Crystallographic studiesof cytochrome c oxidase show an unusual post-translational modification, linking C6 of Tyr(244) and the ε-N of His(240) (bovine enzyme numbering). It plays a vital role in enabling the cytochrome a3- CuBbinuclear center to accept four electrons in reducing molecular oxygen and four protons to water. The mechanism of reduction was formerly thought to involve aperoxideintermediate, which was believed to lead tosuperoxideproduction. However, the currently accepted mechanism involves a rapid four-electron reduction involving immediate oxygen–oxygen bond cleavage, avoiding any intermediate likely to form superoxide.[4]: 865–866
The conserved subunits
[edit]| No. | Subunit name | Humanprotein | Protein description fromUniProt | Pfamfamily with Human protein |
|---|---|---|---|---|
| 1 | Cox1 | COX1_HUMAN | Cytochrome c oxidase subunit 1 | PfamPF00115 |
| 2 | Cox2 | COX2_HUMAN | Cytochrome c oxidase subunit 2 | PfamPF02790,PfamPF00116 |
| 3 | Cox3 | COX3_HUMAN | Cytochrome c oxidase subunit 3 | PfamPF00510 |
| 4 | Cox4i1 | COX41_HUMAN | Cytochrome c oxidase subunit 4 isoform 1, mitochondrial | PfamPF02936 |
| 5 | Cox4a2 | COX42_HUMAN | Cytochrome c oxidase subunit 4 isoform 2, mitochondrial | PfamPF02936 |
| 6 | Cox5a | COX5A_HUMAN | Cytochrome c oxidase subunit 5A, mitochondrial | PfamPF02284 |
| 7 | Cox5b | COX5B_HUMAN | Cytochrome c oxidase subunit 5B, mitochondrial | PfamPF01215 |
| 8 | Cox6a1 | CX6A1_HUMAN | Cytochrome c oxidase subunit 6A1, mitochondrial | PfamPF02046 |
| 9 | Cox6a2 | CX6A2_HUMAN | Cytochrome c oxidase subunit 6A2, mitochondrial | PfamPF02046 |
| 10 | Cox6b1 | CX6B1_HUMAN | Cytochrome c oxidase subunit 6B1 | PfamPF02297 |
| 11 | Cox6b2 | CX6B2_HUMAN | Cytochrome c oxidase subunit 6B2 | PfamPF02297 |
| 12 | Cox6c | COX6C_HUMAN | Cytochrome c oxidase subunit 6C | PfamPF02937 |
| 13 | Cox7a1 | CX7A1_HUMAN | Cytochrome c oxidase subunit 7A1, mitochondrial | PfamPF02238 |
| 14 | Cox7a2 | CX7A2_HUMAN | Cytochrome c oxidase subunit 7A2, mitochondrial | PfamPF02238 |
| 15 | Cox7a3 | COX7S_HUMAN | Putative cytochrome c oxidase subunit 7A3, mitochondrial | PfamPF02238 |
| 16 | Cox7b | COX7B_HUMAN | Cytochrome c oxidase subunit 7B, mitochondrial | PfamPF05392 |
| 17 | Cox7c | COX7C_HUMAN | Cytochrome c oxidase subunit 7C, mitochondrial | PfamPF02935 |
| 18 | Cox7r | COX7R_HUMAN | Cytochrome c oxidase subunit 7A-related protein, mitochondrial | PfamPF02238 |
| 19 | Cox8a | COX8A_HUMAN | Cytochrome c oxidase subunit 8A, mitochondrial P | PfamPF02285 |
| 20 | Cox8c | COX8C_HUMAN | Cytochrome c oxidase subunit 8C, mitochondrial | PfamPF02285 |
| Assembly subunits[7][8][9] | ||||
| 1 | Coa1 | COA1_HUMAN | Cytochrome c oxidase assembly factor 1 homolog | PfamPF08695 |
| 2 | Coa3 | COA3_HUMAN | Cytochrome c oxidase assembly factor 3 homolog, mitochondrial | PfamPF09813 |
| 3 | Coa4 | COA4_HUMAN | Cytochrome c oxidase assembly factor 4 homolog, mitochondrial | PfamPF06747 |
| 4 | Coa5 | COA5_HUMAN | Cytochrome c oxidase assembly factor 5 | PfamPF10203 |
| 5 | Coa6 | COA6_HUMAN | Cytochrome c oxidase assembly factor 6 homolog | PfamPF02297 |
| 6 | Coa7 | COA7_HUMAN | Cytochrome c oxidase assembly factor 7, | PfamPF08238 |
| 7 | Cox11 | COX11_HUMAN | Cytochrome c oxidase assembly protein COX11 mitochondrial | PfamPF04442 |
| 8 | Cox14 | COX14_HUMAN | Cytochrome c oxidase assembly protein | PfamPF14880 |
| 9 | Cox15 | COX15_HUMAN | Cytochrome c oxidase assembly protein COX15 homolog | PfamPF02628 |
| 10 | Cox16 | COX16_HUMAN | Cytochrome c oxidase assembly protein COX16 homolog mitochondrial | PfamPF14138 |
| 11 | Cox17 | COX17_HUMAN | Cytochrome c oxidase copper chaperone | PfamPF05051 |
| 12 | Cox18[10] | COX18_HUMAN | Mitochondrial inner membrane protein (Cytochrome c oxidase assembly protein 18) | PfamPF02096 |
| 13 | Cox19 | COX19_HUMAN | Cytochrome c oxidase assembly protein | PfamPF06747 |
| 14 | Cox20 | COX20_HUMAN | Cytochrome c oxidase protein 20 homolog | PfamPF12597 |
Assembly
[edit]COX assembly inyeastare a complex process that is not entirely understood due to the rapid and irreversible aggregation of hydrophobic subunits that form the holoenzyme complex, as well as aggregation of mutant subunits with exposed hydrophobic patches.[11]COX subunits are encoded in both the nuclear and mitochondrial genomes. The three subunits that form the COX catalytic core are encoded in the mitochondrial genome. Over 30 different nuclear-encoded chaperone proteins are required for COX assembly.[12]
Cofactors, including hemes, are inserted into subunits I & II. The two heme molecules reside in subunit I, helping with transport to subunit II where two copper molecules aid with the continued transfer of electrons.[13]Subunits I and IV initiate assembly. Different subunits may associate to form sub-complex intermediates that later bind to other subunits to form the COX complex.[11]In post-assembly modifications, COX will form a homodimer. This is required for activity. Dimers are connected by acardiolipinmolecule,[11][14][15]which has been found to play a key role in stabilization of the holoenzyme complex. The dissociation of subunits VIIa and III in conjunction with the removal of cardiolipin results in total loss of enzyme activity.[15]Subunits encoded in the nuclear genome are known to play a role in enzyme dimerization and stability. Mutations to these subunits eliminate COX function.[11]
Assembly is known to occur in at least three distinct rate-determining steps. The products of these steps have been found, though specific subunit compositions have not been determined.[11]
Synthesis and assembly of COX subunits I, II, and III are facilitated by translational activators, which interact with the 5’ untranslated regions of mitochondrial mRNA transcripts. Translational activators are encoded in the nucleus. They can operate through either direct or indirect interaction with other components of translation machinery, but exact molecular mechanisms are unclear due to difficulties associated with synthesizing translation machinery in-vitro.[16][17]Though the interactions between subunits I, II, and III encoded within the mitochondrial genome make a lesser contribution to enzyme stability than interactions between bigenomic subunits, these subunits are more conserved, indicating potential unexplored roles for enzyme activity.[18]
Biochemistry
[edit]This sectionis missing informationabout names of the six traditional intermediate states (APFOER); 2021 Cyro-EM result proposing an RPFOE mechanism with reversed assignment of red-ox phases (doi:10.1038/s41467-021-27174-y |
The overall reaction is
- 4 Fe2+– cytochromec+ 4 H++ O2→ 4 Fe3+– cytochromec+ 2 H2O ΔGo' = - 218 kJ/mol,Eo' = +565 mV
Two electrons are passed from two cytochrome c's, through the CuAand cytochrome a sites to the cytochrome a3–CuBbinuclear center, reducing the metals to the Fe2+form and Cu+.The hydroxide ligand is protonated and lost as water, creating a void between the metals that is filled by O2.The oxygen is rapidly reduced, with two electrons coming from the Fe2+-cytochrome a3,which is converted to the ferryl oxo form (Fe4+=O). The oxygen atom close to CuBpicks up one electron from Cu+,and a second electron and a proton from thehydroxylof Tyr(244), which becomes a tyrosyl radical. The second oxygen is converted to a hydroxide ion by picking up two electrons and a proton. A third electron from another cytochrome c is passed through the first two electron carriers to the cytochrome a3–CuBbinuclear center, and this electron and two protons convert the tyrosyl radical back to Tyr, and the hydroxide bound to CuB2+to a water molecule. The fourth electron from another cytochrome c flows through CuAand cytochrome a to the cytochrome a3–CuBbinuclear center, reducing the Fe4+=O to Fe3+,with the oxygen atom picking up a proton simultaneously, regenerating this oxygen as a hydroxide ion coordinated in the middle of the cytochrome a3–CuBcenter as it was at the start of this cycle. Overall, four reduced cytochrome c's are oxidized while O2and four protons are reduced to two water molecules.[4]: 841–5
Inhibition
[edit]COX exists in three conformational states: fully oxidized (pulsed), partially reduced, and fully reduced. Each inhibitor has a high affinity to a different state. In the pulsed state, both the heme a3and the CuBnuclear centers are oxidized; this is the conformation of the enzyme that has the highest activity. A two-electron reduction initiates a conformational change that allows oxygen to bind at the active site to the partially-reduced enzyme. Four electrons bind to COX to fully reduce the enzyme. Its fully reduced state, which consists of a reduced Fe2+at the cytochrome a3heme group and a reduced CuB+binuclear center, is considered the inactive or resting state of the enzyme.[19]
Cyanide,azide,andcarbon monoxide[20]all bind to cytochrome c oxidase, inhibiting the protein from functioning and leading to the chemicalasphyxiationof cells. Higher concentrations of molecular oxygen are needed to compensate for increasing inhibitor concentrations, leading to an overall decrease in metabolic activity in the cell in the presence of an inhibitor. Other ligands, such as nitric oxide and hydrogen sulfide, can also inhibit COX by binding to regulatory sites on the enzyme, reducing the rate of cellular respiration.[21]
Cyanide is a non-competitive inhibitor for COX,[22][23]binding with high affinity to the partially-reduced state of the enzyme and hindering further reduction of the enzyme. In the pulsed state, cyanide binds slowly, but with high affinity. The ligand is posited to electrostatically stabilize both metals at once by positioning itself between them. A high nitric oxide concentration, such as one added exogenously to the enzyme, reverses cyanide inhibition of COX.[24]
Nitric oxidecan reversibly[25]bind to either metal ion in the binuclear center to be oxidized to nitrite. NO and CN−will compete with oxygen to bind at the site, reducing the rate of cellular respiration. Endogenous NO, however, which is produced at lower levels, augments CN−inhibition. Higher levels of NO, which correlate with the existence of more enzyme in the reduced state, lead to a greater inhibition of cyanide.[19]At these basal concentrations, NO inhibition of Complex IV is known to have beneficial effects, such as increasing oxygen levels in blood vessel tissues. The inability of the enzyme to reduce oxygen to water results in a buildup of oxygen, which can diffuse deeper into surrounding tissues.[25]NO inhibition of Complex IV has a larger effect at lower oxygen concentrations, increasing its utility as a vasodilator in tissues of need.[25]
Hydrogen sulfidewill bind COX in a noncompetitive fashion at a regulatory site on the enzyme, similar to carbon monoxide. Sulfide has the highest affinity to either the pulsed or partially reduced states of the enzyme, and is capable of partially reducing the enzyme at the heme a3center. It is unclear whether endogenous H2S levels are sufficient to inhibit the enzyme. There is no interaction between hydrogen sulfide and the fully reduced conformation of COX.[21]
Methanolinmethylated spiritsis converted intoformic acid,which also inhibits the same oxidase system. High levels of ATP canallostericallyinhibit cytochrome c oxidase, binding from within the mitochondrial matrix.[26]
Extramitochondrial and subcellular localizations
[edit]
Cytochrome c oxidase has 3 subunits which are encoded bymitochondrial DNA(cytochrome c oxidasesubunit I,subunit II,andsubunit III). Of these 3 subunits encoded by mitochondrial DNA, two have been identified in extramitochondrial locations. Inpancreaticacinar tissue, these subunits were found inzymogengranules. Additionally, in theanterior pituitary,relatively high amounts of these subunits were found ingrowth hormonesecretory granules.[27]The extramitochondrial function of these cytochrome c oxidase subunits has not yet been characterized. Besides cytochrome c oxidase subunits, extramitochondrial localization has also been observed for large numbers of other mitochondrial proteins.[28][29]This raises the possibility about existence of yet unidentified specific mechanisms for protein translocation from mitochondria to other cellular destinations.[27][29][30]
Genetic defects and disorders
[edit]Defects involving genetic mutations altering cytochromecoxidase (COX) functionality or structure can result in severe, often fatalmetabolic disorders.Such disorders usually manifest in early childhood and affect predominantly tissues with high energy demands (brain, heart, muscle). Among the many classifiedmitochondrial diseases,those involving dysfunctional COX assembly are thought to be the most severe.[31]
The vast majority of COX disorders are linked to mutations in nuclear-encoded proteins referred to as assembly factors, or assembly proteins. These assembly factors contribute to COX structure and functionality, and are involved in several essential processes, including transcription and translation of mitochondrion-encoded subunits, processing of preproteins and membrane insertion, and cofactor biosynthesis and incorporation.[32]
Currently, mutations have been identified in seven COX assembly factors:SURF1,SCO1,SCO2,COX10,COX15,COX20,COA5andLRPPRC.Mutations in these proteins can result in altered functionality of sub-complex assembly, copper transport, or translational regulation. Each gene mutation is associated with the etiology of a specific disease, with some having implications in multiple disorders. Disorders involving dysfunctional COX assembly via gene mutations includeLeigh syndrome,cardiomyopathy,leukodystrophy,anemia,andsensorineural deafness.
Histochemistry
[edit]The increased reliance of neurons on oxidative phosphorylation for energy[33]facilitates the use of COX histochemistry in mapping regional brain metabolism in animals, since it establishes a direct and positive correlation between enzyme activity and neuronal activity.[34]This can be seen in the correlation between COX enzyme amount and activity, which indicates the regulation of COX at the level of gene expression. COX distribution is inconsistent across different regions of the animal brain, but its pattern of its distribution is consistent across animals. This pattern has been observed in the monkey, mouse, and calf brain. One isozyme of COX has been consistently detected in histochemical analysis of the brain.[35]Such brain mapping has been accomplished in spontaneous mutant mice with cerebellar disease such asreeler[36]and a transgenic model ofAlzheimer's disease.[37]This technique has also been used to map learning activity in the animal brain.[38]
Additional images
[edit]-
ETC
-
Complex IV
See also
[edit]- Cytochrome c oxidase subunit I
- Cytochrome c oxidase subunit II
- Cytochrome c oxidase subunit III
- Heme a
References
[edit]- ^Castresana J, Lübben M, Saraste M, Higgins DG (June 1994)."Evolution of cytochrome oxidase, an enzyme older than atmospheric oxygen".The EMBO Journal.13(11): 2516–2525.doi:10.1002/j.1460-2075.1994.tb06541.x.PMC395125.PMID8013452.
- ^Balsa E, Marco R, Perales-Clemente E, Szklarczyk R, Calvo E, Landázuri MO, Enríquez JA (September 2012)."NDUFA4 is a subunit of complex IV of the mammalian electron transport chain".Cell Metabolism.16(3): 378–86.doi:10.1016/j.cmet.2012.07.015.PMID22902835.
- ^Tsukihara T, Aoyama H, Yamashita E, Tomizaki T, Yamaguchi H, Shinzawa-Itoh K, Nakashima R, Yaono R, Yoshikawa S (August 1995). "Structures of metal sites of oxidized bovine heart cytochrome c oxidase at 2.8 A".Science.269(5227): 1069–74.Bibcode:1995Sci...269.1069T.doi:10.1126/science.7652554.PMID7652554.S2CID27210776.
- ^abVoet D, Voet JG (2011).Biochemistry(4th ed.). Hoboken, NJ: John Wiley & Sons.ISBN978-0-470-57095-1.
- ^Zhang Z, Huang L, Shulmeister VM, Chi YI, Kim KK, Hung LW, Crofts AR, Berry EA, Kim SH (April 1998). "Electron transfer by domain movement in cytochrome bc1".Nature.392(6677): 677–84.Bibcode:1998Natur.392..677Z.doi:10.1038/33612.PMID9565029.S2CID4380033.
- ^Kaila VR, Oksanen E, Goldman A, Bloch DA, Verkhovsky MI, Sundholm D, Wikström M (July 2011)."A combined quantum chemical and crystallographic study on the oxidized binuclear center of cytochrome c oxidase".Biochimica et Biophysica Acta (BBA) - Bioenergetics.1807(7): 769–78.doi:10.1016/j.bbabio.2010.12.016.PMID21211513.
- ^Szklarczyk R, Wanschers BF, Cuypers TD, Esseling JJ, Riemersma M, van den Brand MA, Gloerich J, Lasonder E, van den Heuvel LP, Nijtmans LG, Huynen MA (February 2012)."Iterative orthology prediction uncovers new mitochondrial proteins and identifies C12orf62 as the human ortholog of COX14, a protein involved in the assembly of cytochrome c oxidase".Genome Biology.13(2): R12.doi:10.1186/gb-2012-13-2-r12.PMC3334569.PMID22356826.
- ^Mick DU, Dennerlein S, Wiese H, Reinhold R, Pacheu-Grau D, Lorenzi I, Sasarman F, Weraarpachai W, Shoubridge EA, Warscheid B, Rehling P (December 2012)."MITRAC links mitochondrial protein translocation to respiratory-chain assembly and translational regulation".Cell.151(7): 1528–41.doi:10.1016/j.cell.2012.11.053.hdl:11858/00-001M-0000-000E-DDDF-4.PMID23260140.
- ^Kozjak-Pavlovic V, Prell F, Thiede B, Götz M, Wosiek D, Ott C, Rudel T (February 2014). "C1orf163/RESA1 is a novel mitochondrial intermembrane space protein connected to respiratory chain assembly".Journal of Molecular Biology.426(4): 908–20.doi:10.1016/j.jmb.2013.12.001.PMID24333015.
- ^Gaisne M, Bonnefoy N (September 2006)."The COX18 gene, involved in mitochondrial biogenesis, is functionally conserved and tightly regulated in humans and fission yeast".FEMS Yeast Research.6(6): 869–82.doi:10.1111/j.1567-1364.2006.00083.x.PMID16911509.
- ^abcdeFontanesi F, Soto IC, Horn D, Barrientos A (December 2006). "Assembly of mitochondrial cytochrome c-oxidase, a complicated and highly regulated cellular process".American Journal of Physiology. Cell Physiology.291(6): C1129-47.doi:10.1152/ajpcell.00233.2006.PMID16760263.
- ^Dickinson, Elizabeth K.; Adams, Denise L.; Schon, Eric A.; Glerum, D. Moira (September 2000)."A Human SCO2 Mutation Helps Define the Role of Sco1p in the Cytochrome Oxidase Assembly Pathway".Journal of Biological Chemistry.275(35): 26780–26785.doi:10.1016/S0021-9258(19)61443-2.PMID10854440.
- ^Crofts A (1996)."Cytochrome oxidase: Complex IV".University of Illinois at Urbana-Champaign.Archivedfrom the original on 2018-01-23.Retrieved2018-01-28.
- ^Khalimonchuk O, Rödel G (December 2005). "Biogenesis of cytochrome c oxidase".Mitochondrion.5(6): 363–88.doi:10.1016/j.mito.2005.08.002.PMID16199211.
- ^abSedlák E, Robinson NC (September 2015). "Destabilization of the Quaternary Structure of Bovine Heart Cytochrome c Oxidase upon Removal of Tightly Bound Cardiolipin".Biochemistry.54(36): 5569–77.doi:10.1021/acs.biochem.5b00540.PMID26284624.
- ^Herrmann JM, Woellhaf MW, Bonnefoy N (February 2013)."Control of protein synthesis in yeast mitochondria: the concept of translational activators".Biochimica et Biophysica Acta (BBA) - Molecular Cell Research.1833(2): 286–94.doi:10.1016/j.bbamcr.2012.03.007.PMID22450032.
- ^Soto IC, Fontanesi F, Liu J, Barrientos A (June 2012)."Biogenesis and assembly of eukaryotic cytochrome c oxidase catalytic core".Biochimica et Biophysica Acta (BBA) - Bioenergetics.1817(6): 883–97.doi:10.1016/j.bbabio.2011.09.005.PMC3262112.PMID21958598.
- ^Aledo JC, Valverde H, Ruíz-Camacho M, Morilla I, López FD (October 2014)."Protein-protein interfaces from cytochrome c oxidase I evolve faster than nonbinding surfaces, yet negative selection is the driving force".Genome Biology and Evolution.6(11): 3064–76.doi:10.1093/gbe/evu240.PMC4255772.PMID25359921.
- ^abLeavesley HB, Li L, Prabhakaran K, Borowitz JL, Isom GE (January 2008)."Interaction of cyanide and nitric oxide with cytochrome c oxidase: implications for acute cyanide toxicity".Toxicological Sciences.101(1): 101–11.doi:10.1093/toxsci/kfm254.PMID17906319.
- ^Alonso JR, Cardellach F, López S, Casademont J, Miró O (September 2003)."Carbon monoxide specifically inhibits cytochrome c oxidase of human mitochondrial respiratory chain".Pharmacology & Toxicology.93(3): 142–6.doi:10.1034/j.1600-0773.2003.930306.x.PMID12969439.
- ^abNicholls P, Marshall DC, Cooper CE, Wilson MT (October 2013). "Sulfide inhibition of and metabolism by cytochrome c oxidase".Biochemical Society Transactions.41(5): 1312–6.doi:10.1042/BST20130070.PMID24059525.S2CID11554252.
- ^Roberts M, Reiss MJ, Monger G (2000).Advanced Biology.Nelson Thornes.ISBN9780174387329.Archivedfrom the original on 2022-02-24.Retrieved2020-10-25.
- ^Roberts MB (1986).Biology: A Functional Approach.Nelson Thornes.ISBN9780174480198.Archivedfrom the original on 2022-02-24.Retrieved2020-10-25.
- ^Jensen P, Wilson MT, Aasa R, Malmström BG (December 1984)."Cyanide inhibition of cytochrome c oxidase. A rapid-freeze e.p.r. investigation".The Biochemical Journal.224(3): 829–37.doi:10.1042/bj2240829.PMC1144519.PMID6098268.
- ^abcGladwin MT, Shiva S (May 2009)."The ligand binding battle at cytochrome c oxidase: how NO regulates oxygen gradients in tissue".Circulation Research.104(10): 1136–8.doi:10.1161/CIRCRESAHA.109.198911.PMID19461104.
- ^Arnold S, Kadenbach B (October 1997)."Cell respiration s controlled by ATP, an allosteric inhibitor of cytochrome-c oxidase".Eur J Biochem.249(1): 350–354.doi:10.1111/j.1432-1033.1997.t01-1-00350.x.PMID9363790.
- ^abSadacharan SK, Singh B, Bowes T, Gupta RS (November 2005). "Localization of mitochondrial DNA encoded cytochrome c oxidase subunits I and II in rat pancreatic zymogen granules and pituitary growth hormone granules".Histochemistry and Cell Biology.124(5): 409–21.doi:10.1007/s00418-005-0056-2.PMID16133117.S2CID24440427.
- ^Gupta RS, Ramachandra NB, Bowes T, Singh B (2008). "Unusual Cellular Disposition of the Mitochondrial Molecular Chaperones Hsp60, Hsp70 and Hsp10".The Biology of Extracellular Molecular Chaperones.Novartis Foundation Symposia. Vol. 291. pp. 59–68, discussion 69–73, 137–40.doi:10.1002/9780470754030.ch5.ISBN9780470754030.PMID18575266.
{{cite book}}:|journal=ignored (help) - ^abSoltys BJ, Gupta RS (1999). "Mitochondrial proteins at unexpected cellular locations: export of proteins from mitochondria from an evolutionary perspective".International Review of Cytology.194:133–96.doi:10.1016/S0074-7696(08)62396-7.ISBN9780123645982.PMID10494626.
- ^Soltys BJ, Gupta RS (May 1999). "Mitochondrial-matrix proteins at unexpected locations: are they exported?".Trends in Biochemical Sciences.24(5): 174–7.doi:10.1016/s0968-0004(99)01390-0.PMID10322429.
- ^Pecina P, Houstková H, Hansíková H, Zeman J, Houstek J (2004)."Genetic defects of cytochrome c oxidase assembly"(PDF).Physiological Research.53(Suppl 1): S213-23.doi:10.33549/physiolres.930000.53.S213.PMID15119951.S2CID8119738.Archived(PDF)from the original on 2011-07-18.Retrieved2010-11-17.
- ^Zee JM, Glerum DM (December 2006). "Defects in cytochrome oxidase assembly in humans: lessons from yeast".Biochemistry and Cell Biology.84(6): 859–69.doi:10.1139/o06-201.PMID17215873.
- ^Johar K, Priya A, Dhar S, Liu Q, Wong-Riley MT (November 2013)."Neuron-specific specificity protein 4 bigenomically regulates the transcription of all mitochondria- and nucleus-encoded cytochrome c oxidase subunit genes in neurons".Journal of Neurochemistry.127(4): 496–508.doi:10.1111/jnc.12433.PMC3820366.PMID24032355.
- ^Wong-Riley MT (March 1989). "Cytochrome oxidase: an endogenous metabolic marker for neuronal activity".Trends in Neurosciences.12(3): 94–101.doi:10.1016/0166-2236(89)90165-3.PMID2469224.S2CID42996304.
- ^Hevner RF, Wong-Riley MT (November 1989)."Brain cytochrome oxidase: purification, antibody production, and immunohistochemical/histochemical correlations in the CNS".The Journal of Neuroscience.9(11): 3884–98.doi:10.1523/jneurosci.09-11-03884.1989.PMC6569932.PMID2555458.
- ^Strazielle C, Hayzoun K, Derer M, Mariani J, Lalonde R (April 2006). "Regional brain variations of cytochrome oxidase activity in Relnrl-orl mutant mice".Journal of Neuroscience Research.83(5): 821–31.doi:10.1002/jnr.20772.PMID16511878.S2CID45787322.
- ^Strazielle C, Sturchler-Pierrat C, Staufenbiel M, Lalonde R (2003). "Regional brain cytochrome oxidase activity in beta-amyloid precursor protein transgenic mice with the Swedish mutation".Neuroscience.118(4): 1151–63.doi:10.1016/S0306-4522(03)00037-X.PMID12732258.S2CID9366458.
- ^Conejo NM, González-Pardo H, Gonzalez-Lima F, Arias JL (March 2010). "Spatial learning of the water maze: progression of brain circuits mapped with cytochrome oxidase histochemistry".Neurobiology of Learning and Memory.93(3): 362–71.doi:10.1016/j.nlm.2009.12.002.PMID19969098.S2CID24271956.
External links
[edit]- The Cytochrome Oxidase home pageatRice University
- Interactive Molecular model of cytochrome c oxidase(RequiresMDL Chime)
- UMich Orientation of Proteins in Membranesfamilies/superfamily-4
- Cytochrome-c+Oxidaseat the U.S. National Library of MedicineMedical Subject Headings(MeSH)