mTOR
Themammalian target ofrapamycin(mTOR),[5]also referred to as themechanistic target of rapamycin,and sometimes calledFK506-binding protein 12-rapamycin-associated protein 1(FRAP1), is akinasethat in humans is encoded by theMTORgene.[6][7][8]mTOR is a member of thephosphatidylinositol 3-kinase-related kinasefamily ofprotein kinases.[9]
mTOR links with other proteins and serves as a core component of two distinctprotein complexes,mTOR complex 1andmTOR complex 2,which regulate different cellular processes.[10]In particular, as a core component of both complexes, mTOR functions as aserine/threonine protein kinasethat regulates cell growth,cell proliferation,cellmotility,cell survival,protein synthesis,autophagy,andtranscription.[10][11]As a core component of mTORC2, mTOR also functions as atyrosine protein kinasethat promotes the activation ofinsulin receptorsandinsulin-like growth factor 1 receptors.[12]mTORC2 has also been implicated in the control and maintenance of theactin cytoskeleton.[10][13]
Discovery[edit]
Rapa Nui (Easter Island - Chile)[edit]
The study of TOR originated in the 1960s with an expedition toEaster Island(known by the island inhabitants asRapa Nui), with the goal of identifying natural products from plants and soil with possible therapeutic potential. In 1972,Suren Sehgalidentified a small molecule, from the soil bacteriumStreptomyces hygroscopicus,that he purified and initially reported to possess potent antifungal activity. He named itrapamycin,noting its original source and activity.[14][15]Early testing revealed that rapamycin also had potent immunosuppressive and cytostatic anti-cancer activity. Rapamycin did not initially receive significant interest from the pharmaceutical industry until the 1980s, when Wyeth-Ayerst supported Sehgal's efforts to further investigate rapamycin's effect on the immune system. This eventually led to its FDA approval as an immunosuppressant following kidney transplantation. However, prior to its FDA approval, how rapamycin worked remained completely unknown.
Subsequent history[edit]
The discovery of TOR and mTOR stemmed from independent studies of the natural product rapamycin byJoseph Heitman,Rao Movva, andMichael N. Hallin 1991;[16]byDavid M. Sabatini,Hediye Erdjument-Bromage, Mary Lui, Paul Tempst, andSolomon H. Snyderin 1994;[7]and by Candace J. Sabers, Mary M. Martin, Gregory J. Brunn, Josie M. Williams, Francis J. Dumont, Gregory Wiederrecht, and Robert T. Abraham in 1995.[8]In 1991, working in yeast, Hall and colleagues identified the TOR1 and TOR2 genes.[16]In 1993, Robert Cafferkey, George Livi, and colleagues, and Jeannette Kunz,Michael N. Hall,and colleagues independently cloned genes that mediate the toxicity of rapamycin in fungi, known as the TOR/DRR genes.[17][18]
Rapamycin arrests fungal activity at theG1 phaseof the cell cycle. In mammals, it suppresses the immune system by blocking the G1 to S phase transition inT-lymphocytes.[19]Thus, it is used as animmunosuppressantfollowing organ transplantation.[20]Interest in rapamycin was renewed following the discovery of the structurally related immunosuppressive natural productFK506(later called Tacrolimus) in 1987. In 1989–90, FK506 and rapamycin were determined to inhibitT-cell receptor(TCR) andIL-2 receptorsignaling pathways, respectively.[21][22]The two natural products were used to discover theFK506- and rapamycin-binding proteins,includingFKBP12,and to provide evidence that FKBP12–FK506 and FKBP12–rapamycin might act through gain-of-function mechanisms that target distinct cellular functions. These investigations included key studies by Francis Dumont and Nolan Sigal at Merck contributing to show that FK506 and rapamycin behave as reciprocal antagonists.[23][24]These studies implicated FKBP12 as a possible target of rapamycin, but suggested that the complex might interact with another element of the mechanistic cascade.[25][26]
In 1991,calcineurinwas identified as the target of FKBP12-FK506.[27]That of FKBP12-rapamycin remained mysterious until genetic and molecular studies in yeast established FKBP12 as the target of rapamycin, and implicated TOR1 and TOR2 as the targets of FKBP12-rapamycin in 1991 and 1993,[16][28]followed by studies in 1994 when several groups, working independently, discovered the mTOR kinase as its direct target in mammalian tissues.[6][7][20]Sequence analysis of mTOR revealed it to be the direct ortholog of proteins encoded by the yeasttarget of rapamycin 1 and 2 (TOR1 and TOR2) genes, which Joseph Heitman, Rao Movva, andMichael N. Hallhad identified in August 1991 and May 1993. Independently, George Livi and colleagues later reported the same genes, which they calleddominant rapamycin resistance 1 and 2 (DRR1 and DRR2),in studies published in October 1993.
The protein, now called mTOR, was originally named FRAP by Stuart L. Schreiber and RAFT1 by David M. Sabatini;[6][7]FRAP1was used as its official gene symbol in humans. Because of these different names, mTOR, which had been first used by Robert T. Abraham,[6]was increasingly adopted by the community of scientists working on the mTOR pathway to refer to the protein and in homage to the original discovery of the TOR protein in yeast that was named TOR, the Target of Rapamycin, by Joe Heitman, Rao Movva, and Mike Hall. TOR was originally discovered at the Biozentrum and Sandoz Pharmaceuticals in 1991 in Basel, Switzerland, and the name TOR pays further homage to this discovery, as TOR means doorway or gate in German, and the city of Basel was once ringed by a wall punctuated with gates into the city, including the iconicSpalentor.[29]"mTOR" initially meant "mammalian target of rapamycin", but the meaning of the "m" was later changed to "mechanistic".[30]Similarly, with subsequent discoveries the zebra fish TOR was named zTOR, the Arabidopsis thaliana TOR was named AtTOR, and the Drosophila TOR was named dTOR. In 2009 the FRAP1 gene name was officially changed by theHUGO Gene Nomenclature Committee(HGNC) to mTOR, which stands for mechanistic target of rapamycin.[31]
The discovery of TOR and the subsequent identification of mTOR opened the door to the molecular and physiological study of what is now called the mTOR pathway and had a catalytic effect on the growth of the field of chemical biology, where small molecules are used as probes of biology.
Function[edit]
mTOR integrates the input from upstreampathways,includinginsulin,growth factors(such asIGF-1andIGF-2), andamino acids.[11]mTOR also senses cellular nutrient, oxygen, and energy levels.[32]The mTOR pathway is a central regulator of mammalian metabolism and physiology, with important roles in the function of tissues including liver, muscle, white and brown adipose tissue,[33]and the brain, and is dysregulated in human diseases, such asdiabetes,obesity,depression,and certaincancers.[34][35]Rapamycininhibits mTOR by associating with its intracellular receptorFKBP12.[36][37]The FKBP12–rapamycincomplex binds directly to the FKBP12-Rapamycin Binding (FRB) domain of mTOR, inhibiting its activity.[37]
In plants[edit]
Plants express the mechanistic target of rapamycin (mTOR) and have a TOR kinase complex. In plants, only the TORC1 complex is present unlike that of mammalian target of rapamycin which also contains the TORC2 complex.[38]Plant species have TOR proteins in the protein kinase and FKBP-rapamycin binding (FRB) domains that share a similar amino acid sequence to mTOR in mammals.[39]
Role of mTOR in plants
The TOR kinase complex has been known for having a role in the metabolism of plants. The TORC1 complex turns on when plants are living the proper environmental conditions to survive. Once activated, plant cells undergo particular anabolic reactions. These include plant development, translation of mRNA and the growth of cells within the plant. However, the TORC1 complex activation stops catabolic processes such as autophagy from occurring.[38]TOR kinase signaling in plants has been found to aid in senescence, flowering, root and leaf growth, embryogenesis, and the meristem activation above the root cap of a plant.[40]mTOR is also found to be highly involved in developing embryo tissue in plants.[39]
Complexes[edit]

mTOR is thecatalyticsubunit of two structurally distinct complexes: mTORC1 and mTORC2.[41]The two complexes localize to different subcellular compartments, thus affecting their activation and function.[42]Upon activation by Rheb, mTORC1 localizes to theRagulator-Rag complexon the lysosome surface where it then becomes active in the presence of sufficient amino acids.[43][44]
mTORC1[edit]
mTOR Complex 1 (mTORC1) is composed of mTOR, regulatory-associated protein of mTOR (Raptor), mammalian lethal with SEC13 protein 8 (mLST8) and the non-core componentsPRAS40andDEPTOR.[45][46]This complex functions as a nutrient/energy/redox sensor and controls protein synthesis.[11][45]The activity of mTORC1 is regulated byrapamycin,insulin, growth factors,phosphatidic acid,certainamino acidsand their derivatives (e.g.,L-leucineandβ-hydroxy β-methylbutyric acid), mechanical stimuli, andoxidative stress.[45][47][48]
mTORC2[edit]
mTOR Complex 2 (mTORC2) is composed of MTOR, rapamycin-insensitive companion of MTOR (RICTOR),MLST8,and mammalian stress-activated protein kinase interacting protein 1 (mSIN1).[49][50]mTORC2 has been shown to function as an important regulator of theactin cytoskeletonthrough its stimulation of F-actinstress fibers,paxillin,RhoA,Rac1,Cdc42,andprotein kinase Cα (PKCα).[50]mTORC2 also phosphorylates the serine/threonine protein kinaseAkt/PKBon serine residue Ser473, thus affecting metabolism and survival.[51]Phosphorylation of Akt's serine residue Ser473 by mTORC2 stimulates Akt phosphorylation on threonine residue Thr308 byPDK1and leads to full Akt activation.[52][53]In addition, mTORC2 exhibitstyrosine protein kinaseactivity and phosphorylates theinsulin-like growth factor 1 receptor(IGF-1R) andinsulin receptor(InsR) on the tyrosine residues Tyr1131/1136 and Tyr1146/1151, respectively, leading to full activation of IGF-IR and InsR.[12]
Inhibition by rapamycin[edit]
Rapamycin (Sirolimus) inhibits mTORC1, resulting in the suppression ofcellular senescence.[54]This appears to provide most of the beneficial effects of the drug (including life-span extension in animal studies). Suppression ofinsulin resistancebysirtuinsaccounts for at least some of this effect.[55]Impairedsirtuin 3leads tomitochondrial dysfunction.[56]
Rapamycin has a more complex effect on mTORC2, inhibiting it only in certain cell types under prolonged exposure. Disruption of mTORC2 produces the diabetic-like symptoms of decreased glucose tolerance and insensitivity to insulin.[57]
Gene deletion experiments[edit]
The mTORC2 signaling pathway is less defined than the mTORC1 signaling pathway. The functions of the components of the mTORC complexes have been studied usingknockdownsandknockoutsand were found to produce the following phenotypes:
- NIP7:Knockdown reduced mTORC2 activity that is indicated by decreased phosphorylation of mTORC2 substrates.[58]
- RICTOR:Overexpression leads to metastasis and knockdown inhibits growth factor-induced PKC-phosphorylation.[59]Constitutive deletion of Rictor in mice leads to embryonic lethality,[60]while tissue specific deletion leads to a variety of phenotypes; a common phenotype of Rictor deletion in liver, white adipose tissue, and pancreatic beta cells is systemic glucose intolerance and insulin resistance in one or more tissues.[57][61][62][63]Decreased Rictor expression in mice decreases male, but not female, lifespan.[64]
- mTOR: Inhibition of mTORC1 and mTORC2 by PP242 [2-(4-Amino-1-isopropyl-1H-pyrazolo[3,4-d]pyrimidin-3-yl)-1H-indol-5-ol] leads toautophagyorapoptosis;inhibition of mTORC2 alone by PP242 prevents phosphorylation of Ser-473 site on AKT and arrests the cells inG1 phaseof thecell cycle.[65]Genetic reduction of mTOR expression in mice significantly increases lifespan.[66]
- PDK1:Knockout is lethal;hypomorphic alleleresults in smaller organ volume and organism size but normal AKT activation.[67]
- AKT:Knockout mice experience spontaneous apoptosis (AKT1), severe diabetes (AKT2), small brains (AKT3), and growth deficiency (AKT1/AKT2).[68]Mice heterozygous for AKT1 have increased lifespan.[69]
- TOR1, theS. cerevisiaeorthologue of mTORC1, is a regulator of both carbon and nitrogen metabolism; TOR1 KO strains regulate response to nitrogen as well as carbon availability, indicating that it is a key nutritional transducer in yeast.[70][71]
Clinical significance[edit]
Aging[edit]

Decreased TOR activity has been found to increase life span inS. cerevisiae,C. elegans,andD. melanogaster.[72][73][74][75]The mTOR inhibitorrapamycinhas been confirmed to increase lifespan in mice.[76][77][78][79][80]
It is hypothesized that some dietary regimes, likecaloric restrictionandmethioninerestriction, cause lifespan extension by decreasing mTOR activity.[72][73]Some studies have suggested that mTOR signaling may increase during aging, at least in specific tissues like adipose tissue, and rapamycin may act in part by blocking this increase.[81]An alternative theory is mTOR signaling is an example ofantagonistic pleiotropy,and while high mTOR signaling is good during early life, it is maintained at an inappropriately high level in old age. Calorie restriction and methionine restriction may act in part by limiting levels ofessential amino acidsincluding leucine and methionine, which are potent activators of mTOR.[82]The administration ofleucineinto the rat brain has been shown to decrease food intake and body weight via activation of the mTOR pathway in thehypothalamus.[83]
According to thefree radical theory of aging,[84]reactive oxygen speciescause damage tomitochondrialproteins and decrease ATP production. Subsequently, via ATP sensitiveAMPK,the mTOR pathway is inhibited and ATP-consuming protein synthesis is downregulated, since mTORC1 initiates a phosphorylation cascade activating theribosome.[19]Hence, the proportion of damaged proteins is enhanced. Moreover, disruption of mTORC1 directly inhibitsmitochondrial respiration.[85]These positive feedbacks on the aging process are counteracted by protective mechanisms: Decreased mTOR activity (among other factors) upregulates removal of dysfunctional cellular components viaautophagy.[84]
mTOR is a key initiator of thesenescence-associated secretory phenotype(SASP).[86]Interleukin 1 alpha(IL1A) is found on the surface ofsenescent cellswhere it contributes to the production of SASP factors due to apositive feedback loopwith NF-κB.[87][88]Translation ofmRNAfor IL1A is highly dependent upon mTOR activity.[89]mTOR activity increases levels of IL1A, mediated byMAPKAPK2.[87]mTOR inhibition ofZFP36L1prevents this protein from degradingtranscriptsof numerous components of SASP factors.[90]
Cancer[edit]
Over-activation of mTOR signaling significantly contributes to the initiation and development of tumors and mTOR activity was found to be deregulated in many types of cancer including breast, prostate, lung, melanoma, bladder, brain, and renal carcinomas.[91]Reasons for constitutive activation are several. Among the most common are mutations in tumor suppressorPTENgene. PTEN phosphatase negatively affects mTOR signalling through interfering with the effect ofPI3K,an upstream effector of mTOR. Additionally, mTOR activity is deregulated in many cancers as a result of increased activity of PI3K orAkt.[92]Similarly, overexpression of downstream mTOR effectors4E-BP1,S6K1,S6K2andeIF4Eleads to poor cancer prognosis.[93]Also, mutations inTSCproteins that inhibit the activity of mTOR may lead to a condition namedtuberous sclerosis complex,which exhibits as benign lesions and increases the risk ofrenal cell carcinoma.[94]
Increasing mTOR activity was shown to drive cell cycle progression and increase cell proliferation mainly due to its effect on protein synthesis. Moreover, active mTOR supports tumor growth also indirectly by inhibitingautophagy.[95]Constitutively activated mTOR functions in supplying carcinoma cells with oxygen and nutrients by increasing the translation ofHIF1Aand supportingangiogenesis.[96]mTOR also aids in another metabolic adaptation of cancerous cells to support their increased growth rate—activation ofglycolytic metabolism.Akt2,a substrate of mTOR, specifically ofmTORC2,upregulates expression of the glycolytic enzymePKM2thus contributing to theWarburg effect.[97]
Central nervous system disorders / Brain function[edit]
Autism[edit]
mTOR is implicated in the failure of a 'pruning' mechanism of the excitatory synapses inautism spectrumdisorders.[98]
Alzheimer's disease[edit]
mTOR signaling intersects withAlzheimer's disease(AD) pathology in several aspects, suggesting its potential role as a contributor to disease progression. In general, findings demonstrate mTOR signaling hyperactivity in AD brains. For example, postmortem studies of human AD brain reveal dysregulation in PTEN, Akt, S6K, and mTOR.[99][100][101]mTOR signaling appears to be closely related to the presence of soluble amyloid beta (Aβ) and tau proteins, which aggregate and form two hallmarks of the disease, Aβ plaques and neurofibrillary tangles, respectively.[102]In vitro studies have shown Aβ to be an activator of thePI3K/AKT pathway,which in turn activates mTOR.[103]In addition, applying Aβ to N2K cells increases the expression of p70S6K, a downstream target of mTOR known to have higher expression in neurons that eventually develop neurofibrillary tangles.[104][105]Chinese hamster ovary cells transfected with the 7PA2 familial AD mutation also exhibit increased mTOR activity compared to controls, and the hyperactivity is blocked using a gamma-secretase inhibitor.[106][107]These in vitro studies suggest that increasing Aβ concentrations increases mTOR signaling; however, significantly large, cytotoxic Aβ concentrations are thought to decrease mTOR signaling.[108]
Consistent with data observed in vitro, mTOR activity and activated p70S6K have been shown to be significantly increased in the cortex and hippocampus of animal models of AD compared to controls.[107][109]Pharmacologic or genetic removal of the Aβ in animal models of AD eliminates the disruption in normal mTOR activity, pointing to the direct involvement of Aβ in mTOR signaling.[109]In addition, by injecting Aβ oligomers into the hippocampi of normal mice, mTOR hyperactivity is observed.[109]Cognitive impairments characteristic of AD appear to be mediated by the phosphorylation of PRAS-40, which detaches from and allows for the mTOR hyperactivity when it is phosphorylated; inhibiting PRAS-40 phosphorylation prevents Aβ-induced mTOR hyperactivity.[109][110][111]Given these findings, the mTOR signaling pathway appears to be one mechanism of Aβ-induced toxicity in AD.
The hyperphosphorylation of tau proteins into neurofibrillary tangles is one hallmark of AD. p70S6K activation has been shown to promote tangle formation as well as mTOR hyperactivity through increased phosphorylation and reduced dephosphorylation.[104][112][113][114]It has also been proposed that mTOR contributes to tau pathology by increasing the translation of tau and other proteins.[115]
Synaptic plasticity is a key contributor to learning and memory, two processes that are severely impaired in AD patients. Translational control, or the maintenance of protein homeostasis, has been shown to be essential for neural plasticity and is regulated by mTOR.[107][116][117][118][119]Both protein over- and under-production via mTOR activity seem to contribute to impaired learning and memory. Furthermore, given that deficits resulting from mTOR overactivity can be alleviated through treatment with rapamycin, it is possible that mTOR plays an important role in affecting cognitive functioning through synaptic plasticity.[103][120]Further evidence for mTOR activity in neurodegeneration comes from recent findings demonstrating that eIF2α-P, an upstream target of the mTOR pathway, mediates cell death in prion diseases through sustained translational inhibition.[121]
Some evidence points to mTOR's role in reduced Aβ clearance as well. mTOR is a negative regulator of autophagy;[122]therefore, hyperactivity in mTOR signaling should reduce Aβ clearance in the AD brain. Disruptions in autophagy may be a potential source of pathogenesis in protein misfolding diseases, including AD.[123][124][125][126][127][128]Studies using mouse models of Huntington's disease demonstrate that treatment with rapamycin facilitates the clearance of huntingtin aggregates.[129][130]Perhaps the same treatment may be useful in clearing Aβ deposits as well.
Lymphoproliferative diseases[edit]
Hyperactive mTOR pathways have been identified in certain lymphoproliferative diseases such asautoimmune lymphoproliferative syndrome(ALPS),[131]multicentricCastleman disease,[132]andpost-transplant lymphoproliferative disorder(PTLD).[133]
Protein synthesis and cell growth[edit]
mTORC1 activation is required for myofibrillar muscle protein synthesis and skeletalmuscle hypertrophyin humans in response to bothphysical exerciseand ingestion of certainamino acidsor amino acid derivatives.[134][135]Persistent inactivation of mTORC1 signaling in skeletal muscle facilitates the loss of muscle mass and strength during muscle wasting in old age,cancer cachexia,andmuscle atrophyfromphysical inactivity.[134][135][136]mTORC2 activation appears to mediateneuriteoutgrowth in differentiated mouseneuro2a cells.[137]Intermittent mTOR activation inprefrontalneurons byβ-hydroxy β-methylbutyrateinhibits age-related cognitive decline associated with dendritic pruning in animals, which is a phenomenon also observed in humans.[138]
• PA:phosphatidic acid
• mTOR: mechanistic target of rapamycin
• AMP:adenosine monophosphate
• ATP:adenosine triphosphate
• AMPK:AMP-activated protein kinase
• PGC‐1α:peroxisome proliferator-activated receptor gamma coactivator-1α
• S6K1:p70S6 kinase
• 4EBP1:eukaryotic translation initiation factor 4E-binding protein 1
• eIF4E:eukaryotic translation initiation factor 4E
• RPS6:ribosomal protein S6
• eEF2:eukaryotic elongation factor 2
• RE:resistance exercise;EE:endurance exercise
• Myo:myofibrillar;Mito:mitochondrial
• AA:amino acids
• HMB:β-hydroxy β-methylbutyric acid
• ↑ represents activation
• Τ represents inhibition
Lysosomal damage inhibits mTOR and induces autophagy[edit]
ActivemTORC1is positioned onlysosomes.mTORis inhibited[140]when lysosomal membrane is damaged by various exogenous or endogenous agents, such as invadingbacteria,membrane-permeant chemicals yielding osmotically active products (this type of injury can be modeled using membrane-permeant dipeptide precursors that polymerize in lysosomes),amyloidprotein aggregates(see above section onAlzheimer's disease) and cytoplasmic organic or inorganicinclusionsincludinguratecrystals andcrystalline silica.[140]The process of mTOR inactivation following lysosomal/endomembrane is mediated by the protein complex termed GALTOR.[140]At the heart of GALTOR[140]isgalectin-8,a member of β-galactoside binding superfamily of cytosolic lectins termedgalectins,which recognizes lysosomal membrane damage by binding to the exposedglycanson the lumenal side of the delimiting endomembrane. Following membrane damage, galectin-8, which normally associates with mTOR under homeostatic conditions, no longer interacts with mTOR but now instead binds toSLC38A9,RRAGA/RRAGB,andLAMTOR1,inhibitingRagulator's (LAMTOR1-5 complex)guanine nucleotide exchangefunction-[140]
TORis a negative regulator of autophagy in general, best studied during response to starvation,[141][142][143][144][145]which is a metabolic response. During lysosomal damage however, mTOR inhibition activatesautophagyresponse in its quality control function, leading to the process termed lysophagy[146]that removes damaged lysosomes. At this stage anothergalectin,galectin-3,interacts withTRIM16to guide selective autophagy of damaged lysosomes.[147][148]TRIM16 gathersULK1and principal components(Beclin 1andATG16L1) of other complexes (Beclin 1-VPS34-ATG14andATG16L1-ATG5-ATG12) initiatingautophagy,[148]many of them being under negative control of mTOR directly such as the ULK1-ATG13 complex,[143][144][145]or indirectly, such as components of the class III PI3K(Beclin 1, ATG14 and VPS34) since they depend on activating phosphorylations by ULK1 when it is not inhibited by mTOR. Theseautophagy-driving components physically and functionally link up with each other integrating all processes necessary for autophagosomal formation: (i) the ULK1-ATG13-FIP200/RB1CC1complex associates with theLC3B/GABARAPconjugation machinery through direct interactions betweenFIP200/RB1CC1andATG16L1,[149][150][151](ii)ULK1-ATG13-FIP200/RB1CC1complex associates with theBeclin 1-VPS34-ATG14via direct interactions betweenATG13'sHORMA domainandATG14,[152](iii) ATG16L1 interacts withWIPI2,which binds toPI3P,the enzymatic product of the class III PI3K Beclin 1-VPS34-ATG14.[153]Thus, mTOR inactivation, initiated through GALTOR[140]upon lysosomal damage, plus a simultaneous activation viagalectin-9(which also recognizes lysosomal membrane breach) ofAMPK[140]that directly phosphorylates and activates key components (ULK1,[154]Beclin 1[155]) of the autophagy systems listed above and further inactivates mTORC1,[156][157]allows for strong autophagy induction and autophagic removal of damaged lysosomes.
Additionally, several types of ubiquitination events parallel and complement the galectin-driven processes:Ubiquitinationof TRIM16-ULK1-Beclin-1 stabilizes these complexes to promote autophagy activation as described above.[148]ATG16L1has an intrinsic binding affinity forubiquitin[151]); whereas ubiquitination by a glycoprotein-specific FBXO27-endowed ubiquitin ligase of several damage-exposed glycosylated lysosomal membrane proteins such asLAMP1,LAMP2,GNS/N-acetylglucosamine-6-sulfatase,TSPAN6/tetraspanin-6,PSAP/prosaposin,and TMEM192/transmembrane protein 192[158]may contribute to the execution of lysophagy via autophagic receptors such as p62/SQSTM1,which is recruited during lysophagy,[151]or other to be determined functions.
Scleroderma[edit]
Scleroderma,also known as systemicsclerosis,is a chronicsystemic autoimmune diseasecharacterised by hardening (sclero) of the skin (derma) that affects internal organs in its more severe forms.[159][160]mTOR plays a role infibroticdiseases and autoimmunity, and blockade of the mTORC pathway is under investigation as a treatment for scleroderma.[9]
mTOR inhibitors as therapies[edit]
Transplantation[edit]
mTOR inhibitors, e.g.rapamycin,are already used to preventtransplant rejection.
Glycogen storage disease[edit]
Some articles reported that rapamycin can inhibit mTORC1 so that the phosphorylation of GS (glycogen synthase) can be increased in skeletal muscle. This discovery represents a potential novel therapeutic approach forglycogen storage diseasethat involve glycogen accumulation in muscle.
Anti-cancer[edit]
There are two primary mTOR inhibitors used in the treatment of human cancers,temsirolimusandeverolimus.mTOR inhibitors have found use in the treatment of a variety of malignancies, includingrenal cell carcinoma(temsirolimus) andpancreatic cancer,breast cancer,and renal cell carcinoma (everolimus).[161]The complete mechanism of these agents is not clear, but they are thought to function by impairing tumourangiogenesisand causing impairment of theG1/S transition.[162]
Anti-aging[edit]
mTOR inhibitors may be useful for treating/preventing several age-associated conditions,[163]including neurodegenerative diseases such asAlzheimer's diseaseandParkinson's disease.[164]After a short-term treatment with the mTOR inhibitorsdactolisibandeverolimus,in elderly (65 and older), treated subjects had a reduced number of infections over the course of a year.[165]
Various natural compounds, includingepigallocatechin gallate(EGCG),caffeine,curcumin,berberine,quercetin,resveratrolandpterostilbene,have been reported to inhibit mTOR when applied to isolated cells in culture.[166][167][168]As yet no high quality evidence exists that these substances inhibit mTOR signaling or extend lifespan when taken asdietary supplementsby humans, despite encouraging results in animals such as fruit flies and mice. Various trials are ongoing.[169][170]
Interactions[edit]
Mechanistic target of rapamycin has been shown tointeractwith:[171]
- ABL1,[172]
- AKT1,[52][173][174]
- IGF-IR,[12]
- InsR,[12]
- CLIP1,[175]
- EIF3F[176]
- EIF4EBP1,[45][177][178][179][180][181][182][183]
- FKBP1A,[13][50][184][185][186][187]
- GPHN,[188]
- KIAA1303,[13][45][49][50][85][177][178][179][189][190][191][192][193][194][195][196][197][198][199][200]
- PRKCD,[201]
- RHEB,[180][202][203][204]
- RICTOR,[13][49][50][191][197][199][200]
- RPS6KB1,[45][178][180][181][182][196][199][205][206][207][208][209][210][211][212]
- STAT1,[213]
- STAT3,[214][215]
- Two-pore channels:TPCN1;TPCN2,[216]and
- UBQLN1.[217]
References[edit]
- ^abcGRCh38: Ensembl release 89: ENSG00000198793–Ensembl,May 2017
- ^abcGRCm38: Ensembl release 89: ENSMUSG00000028991–Ensembl,May 2017
- ^"Human PubMed Reference:".National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^"Mouse PubMed Reference:".National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^Sabers CJ, Martin MM, Brunn GJ, et al. (Jan 1995)."Isolation of a Protein Target of the FKBP12-Rapamycin Complex in Mammalian Cells".J. Biol. Chem.270(2): 815–22.doi:10.1074/jbc.270.2.815.PMID7822316.
- ^abcdBrown EJ, Albers MW, Shin TB, et al. (June 1994). "A mammalian protein targeted by G1-arresting rapamycin-receptor complex".Nature.369(6483): 756–8.Bibcode:1994Natur.369..756B.doi:10.1038/369756a0.PMID8008069.S2CID4359651.
- ^abcdSabatini DM, Erdjument-Bromage H, Lui M, et al. (July 1994). "RAFT1: a mammalian protein that binds to FKBP12 in a rapamycin-dependent fashion and is homologous to yeast TORs".Cell.78(1): 35–43.doi:10.1016/0092-8674(94)90570-3.PMID7518356.S2CID33647539.
- ^abSabers CJ, Martin MM, Brunn GJ, et al. (January 1995)."Isolation of a protein target of the FKBP12-rapamycin complex in mammalian cells".The Journal of Biological Chemistry.270(2): 815–22.doi:10.1074/jbc.270.2.815.PMID7822316.
- ^abMitra A, Luna JI, Marusina AI, et al. (November 2015)."Dual mTOR Inhibition Is Required to Prevent TGF-β-Mediated Fibrosis: Implications for Scleroderma".The Journal of Investigative Dermatology.135(11): 2873–6.doi:10.1038/jid.2015.252.PMC4640976.PMID26134944.
- ^abcdefLipton JO, Sahin M (October 2014)."The neurology of mTOR".Neuron.84(2): 275–291.doi:10.1016/j.neuron.2014.09.034.PMC4223653.PMID25374355.
The mTOR signaling pathway acts as a molecular systems integrator to support organismal and cellular interactions with the environment. The mTOR pathway regulates homeostasis by directly influencing protein synthesis, transcription, autophagy, metabolism, and organelle biogenesis and maintenance. It is not surprising then that mTOR signaling is implicated in the entire hierarchy of brain function including the proliferation of neural stem cells, the assembly and maintenance of circuits, experience-dependent plasticity and regulation of complex behaviors like feeding, sleep and circadian rhythms....
mTOR function is mediated through two large biochemical complexes defined by their respective protein composition and have been extensively reviewed elsewhere(Dibble and Manning, 2013; Laplante and Sabatini, 2012)(Figure 1B). In brief, common to both mTOR complex 1 (mTORC1) and mTOR complex 2 (mTORC2) are: mTOR itself, mammalian lethal with sec13 protein 8 (mLST8; also known as GβL), and the inhibitory DEP domain containing mTOR-interacting protein (DEPTOR). Specific to mTORC1 is the regulator-associated protein of the mammalian target of rapamycin (Raptor) and proline-rich Akt substrate of 40 kDa (PRAS40)(Kim et al., 2002; Laplante and Sabatini, 2012). Raptor is essential to mTORC1 activity. The mTORC2 complex includes the rapamycin insensitive companion of mTOR (Rictor), mammalian stress activated MAP kinase-interacting protein 1 (mSIN1), and proteins observed with rictor 1 and 2 (PROTOR 1 and 2)(Jacinto et al., 2006; Jacinto et al., 2004; Pearce et al., 2007; Sarbassov et al., 2004)(Figure 1B). Rictor and mSIN1 are both critical to mTORC2 function.
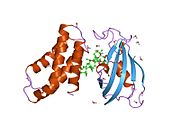
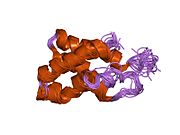
Figure 1: Domain structure of the mTOR kinase and components of mTORC1 and mTORC2
Figure 2: The mTOR Signaling Pathway - ^abcHay N, Sonenberg N (August 2004)."Upstream and downstream of mTOR".Genes & Development.18(16): 1926–45.doi:10.1101/gad.1212704.PMID15314020.
- ^abcdYin Y, Hua H, Li M, et al. (January 2016)."mTORC2 promotes type I insulin-like growth factor receptor and insulin receptor activation through the tyrosine kinase activity of mTOR".Cell Research.26(1): 46–65.doi:10.1038/cr.2015.133.PMC4816127.PMID26584640.
- ^abcdJacinto E, Loewith R, Schmidt A, et al. (November 2004). "Mammalian TOR complex 2 controls the actin cytoskeleton and is rapamycin insensitive".Nature Cell Biology.6(11): 1122–8.doi:10.1038/ncb1183.PMID15467718.S2CID13831153.
- ^Powers T (November 2022). Kellogg D (ed.)."The origin story of rapamycin: systemic bias in biomedical research and cold war politics".Molecular Biology of the Cell.33(13).doi:10.1091/mbc.E22-08-0377.PMC9634974.PMID36228182.
- ^Sehgal SN, Baker H, Vézina C (October 1975). "Rapamycin (AY-22,989), a new antifungal antibiotic. II. Fermentation, isolation and characterization".The Journal of Antibiotics.28(10): 727–732.doi:10.7164/antibiotics.28.727.PMID1102509.
- ^abcHeitman J, Movva NR, Hall MN (August 1991). "Targets for cell cycle arrest by the immunosuppressant rapamycin in yeast".Science.253(5022): 905–9.Bibcode:1991Sci...253..905H.doi:10.1126/science.1715094.PMID1715094.S2CID9937225.
- ^Kunz J, Henriquez R, Schneider U, et al. (May 1993). "Target of rapamycin in yeast, TOR2, is an essential phosphatidylinositol kinase homolog required for G1 progression".Cell.73(3): 585–596.doi:10.1016/0092-8674(93)90144-F.PMID8387896.S2CID42926249.
- ^Cafferkey R, Young PR, McLaughlin MM, et al. (October 1993)."Dominant missense mutations in a novel yeast protein related to mammalian phosphatidylinositol 3-kinase and VPS34 abrogate rapamycin cytotoxicity".Mol Cell Biol.13(10): 6012–23.doi:10.1128/MCB.13.10.6012.PMC364661.PMID8413204.
- ^abMagnuson B, Ekim B, Fingar DC (January 2012). "Regulation and function of ribosomal protein S6 kinase (S6K) within mTOR signaling networks".The Biochemical Journal.441(1): 1–21.doi:10.1042/BJ20110892.PMID22168436.S2CID12932678.
- ^abAbraham RT, Wiederrecht GJ (1996). "Immunopharmacology of rapamycin".Annual Review of Immunology.14:483–510.doi:10.1146/annurev.immunol.14.1.483.PMID8717522.
- ^Bierer BE, Mattila PS, Standaert RF, et al. (December 1990)."Two distinct signal transmission pathways in T lymphocytes are inhibited by complexes formed between an immunophilin and either FK506 or rapamycin".Proceedings of the National Academy of Sciences of the United States of America.87(23): 9231–5.Bibcode:1990PNAS...87.9231B.doi:10.1073/pnas.87.23.9231.PMC55138.PMID2123553.
- ^Bierer BE, Somers PK, Wandless TJ, et al. (October 1990). "Probing immunosuppressant action with a nonnatural immunophilin ligand".Science.250(4980): 556–9.Bibcode:1990Sci...250..556B.doi:10.1126/science.1700475.PMID1700475.S2CID11123023.
- ^Dumont FJ, Melino MR, Staruch MJ, et al. (February 1990)."The immunosuppressive macrolides FK-506 and rapamycin act as reciprocal antagonists in murine T cells".J Immunol.144(4): 1418–24.doi:10.4049/jimmunol.144.4.1418.PMID1689353.S2CID44256944.
- ^Dumont FJ, Staruch MJ, Koprak SL, et al. (January 1990)."Distinct mechanisms of suppression of murine T cell activation by the related macrolides FK-506 and rapamycin".J Immunol.144(1): 251–8.doi:10.4049/jimmunol.144.1.251.PMID1688572.S2CID13201695.
- ^Harding MW, Galat A, Uehling DE, et al. (October 1989). "A receptor for the immunosuppressant FK506 is a cis-trans peptidyl-prolyl isomerase".Nature.341(6244): 758–60.Bibcode:1989Natur.341..758H.doi:10.1038/341758a0.PMID2477715.S2CID4349152.
- ^Fretz H, Albers MW, Galat A, et al. (February 1991). "Rapamycin and FK506 binding proteins (immunophilins)".Journal of the American Chemical Society.113(4): 1409–1411.doi:10.1021/ja00004a051.
- ^Liu J, Farmer JD, Lane WS, et al. (August 1991). "Calcineurin is a common target of cyclophilin-cyclosporin A and FKBP-FK506 complexes".Cell.66(4): 807–15.doi:10.1016/0092-8674(91)90124-H.PMID1715244.S2CID22094672.
- ^Kunz J, Henriquez R, Schneider U, et al. (May 1993). "Target of rapamycin in yeast, TOR2, is an essential phosphatidylinositol kinase homolog required for G1 progression".Cell.73(3): 585–596.doi:10.1016/0092-8674(93)90144-F.PMID8387896.S2CID42926249.
- ^Heitman J (November 2015)."On the discovery of TOR as the target of rapamycin".PLOS Pathogens.11(11): e1005245.doi:10.1371/journal.ppat.1005245.PMC4634758.PMID26540102.
- ^Kennedy BK, Lamming DW (2016)."The Mechanistic Target of Rapamycin: The Grand ConducTOR of Metabolism and Aging".Cell Metabolism.23(6): 990–1003.doi:10.1016/j.cmet.2016.05.009.PMC4910876.PMID27304501.
- ^"Symbol report for MTOR".HGNC data for MTOR.HUGO Gene Nomenclature Committee.September 1, 2020.Retrieved2020-12-17.
- ^Tokunaga C, Yoshino K, Yonezawa K (January 2004). "mTOR integrates amino acid- and energy-sensing pathways".Biochemical and Biophysical Research Communications.313(2): 443–6.doi:10.1016/j.bbrc.2003.07.019.PMID14684182.
- ^Wipperman MF, Montrose DC, Gotto AM, et al. (2019)."Mammalian Target of Rapamycin: A Metabolic Rheostat for Regulating Adipose Tissue Function and Cardiovascular Health".The American Journal of Pathology.189(3): 492–501.doi:10.1016/j.ajpath.2018.11.013.PMC6412382.PMID30803496.
- ^Beevers CS, Li F, Liu L, et al. (August 2006). "Curcumin inhibits the mammalian target of rapamycin-mediated signaling pathways in cancer cells".International Journal of Cancer.119(4): 757–64.doi:10.1002/ijc.21932.PMID16550606.S2CID25454463.
- ^Kennedy BK, Lamming DW (June 2016)."The Mechanistic Target of Rapamycin: The Grand ConducTOR of Metabolism and Aging".Cell Metabolism.23(6): 990–1003.doi:10.1016/j.cmet.2016.05.009.PMC4910876.PMID27304501.
- ^Huang S, Houghton PJ (December 2001). "Mechanisms of resistance to rapamycins".Drug Resistance Updates.4(6): 378–91.doi:10.1054/drup.2002.0227.PMID12030785.
- ^abHuang S, Bjornsti MA, Houghton PJ (2003)."Rapamycins: mechanism of action and cellular resistance".Cancer Biology & Therapy.2(3): 222–32.doi:10.4161/cbt.2.3.360.PMID12878853.
- ^abIngargiola C, Turqueto Duarte G, Robaglia C, et al. (October 2020)."The Plant Target of Rapamycin: A Conduc TOR of Nutrition and Metabolism in Photosynthetic Organisms".Genes.11(11): 1285.doi:10.3390/genes11111285.PMC7694126.PMID33138108.
- ^abShi L, Wu Y, Sheen J (July 2018)."TOR signaling in plants: conservation and innovation".Development.145(13).doi:10.1242/dev.160887.PMC6053665.PMID29986898.
- ^Xiong Y, Sheen J (February 2014)."The role of target of rapamycin signaling networks in plant growth and metabolism".Plant Physiology.164(2): 499–512.doi:10.1104/pp.113.229948.PMC3912084.PMID24385567.
- ^Wullschleger S, Loewith R, Hall MN (February 2006)."TOR signaling in growth and metabolism".Cell.124(3): 471–84.doi:10.1016/j.cell.2006.01.016.PMID16469695.
- ^Betz C, Hall MN (November 2013)."Where is mTOR and what is it doing there?".The Journal of Cell Biology.203(4): 563–74.doi:10.1083/jcb.201306041.PMC3840941.PMID24385483.
- ^Groenewoud MJ, Zwartkruis FJ (August 2013). "Rheb and Rags come together at the lysosome to activate mTORC1".Biochemical Society Transactions.41(4): 951–5.doi:10.1042/bst20130037.PMID23863162.S2CID8237502.
- ^Efeyan A, Zoncu R, Sabatini DM (September 2012)."Amino acids and mTORC1: from lysosomes to disease".Trends in Molecular Medicine.18(9): 524–33.doi:10.1016/j.molmed.2012.05.007.PMC3432651.PMID22749019.
- ^abcdefKim DH, Sarbassov DD, Ali SM, et al. (July 2002)."mTOR interacts with raptor to form a nutrient-sensitive complex that signals to the cell growth machinery".Cell.110(2): 163–75.doi:10.1016/S0092-8674(02)00808-5.PMID12150925.
- ^Kim DH, Sarbassov DD, Ali SM, et al. (April 2003)."GbetaL, a positive regulator of the rapamycin-sensitive pathway required for the nutrient-sensitive interaction between raptor and mTOR".Molecular Cell.11(4): 895–904.doi:10.1016/S1097-2765(03)00114-X.PMID12718876.
- ^Fang Y, Vilella-Bach M, Bachmann R, et al. (November 2001). "Phosphatidic acid-mediated mitogenic activation of mTOR signaling".Science.294(5548): 1942–5.Bibcode:2001Sci...294.1942F.doi:10.1126/science.1066015.PMID11729323.S2CID44444716.
- ^Bond P (March 2016)."Regulation of mTORC1 by growth factors, energy status, amino acids and mechanical stimuli at a glance".J. Int. Soc. Sports Nutr.13:8.doi:10.1186/s12970-016-0118-y.PMC4774173.PMID26937223.
- ^abcFrias MA, Thoreen CC, Jaffe JD, et al. (September 2006)."mSin1 is necessary for Akt/PKB phosphorylation, and its isoforms define three distinct mTORC2s".Current Biology.16(18): 1865–70.Bibcode:2006CBio...16.1865F.doi:10.1016/j.cub.2006.08.001.PMID16919458.
- ^abcdeSarbassov DD, Ali SM, Kim DH, et al. (July 2004)."Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton".Current Biology.14(14): 1296–302.Bibcode:2004CBio...14.1296D.doi:10.1016/j.cub.2004.06.054.PMID15268862.
- ^Betz C, Stracka D, Prescianotto-Baschong C, et al. (July 2013)."Feature Article: mTOR complex 2-Akt signaling at mitochondria-associated endoplasmic reticulum membranes (MAM) regulates mitochondrial physiology".Proceedings of the National Academy of Sciences of the United States of America.110(31): 12526–34.doi:10.1073/pnas.1302455110.PMC3732980.PMID23852728.
- ^abSarbassov DD, Guertin DA, Ali SM, et al. (February 2005). "Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex".Science.307(5712): 1098–101.Bibcode:2005Sci...307.1098S.doi:10.1126/science.1106148.PMID15718470.S2CID45837814.
- ^Stephens L, Anderson K, Stokoe D, et al. (January 1998). "Protein kinase B kinases that mediate phosphatidylinositol 3,4,5-trisphosphate-dependent activation of protein kinase B".Science.279(5351): 710–4.Bibcode:1998Sci...279..710S.doi:10.1126/science.279.5351.710.PMID9445477.
- ^Carosi JM, Fourrier C, Bensalem J, et al. (2022)."The mTOR-lysosome axis at the centre of ageing".FEBS Open Bio.12(4): 739–757.doi:10.1002/2211-5463.13347.PMC8972043.PMID34878722.
- ^Zhou S, Tang X, Chen H (2018)."Sirtuins and Insulin Resistance".Frontiers in Endocrinology.9:748.doi:10.3389/fendo.2018.00748.PMC6291425.PMID30574122.
- ^Baechle JJ, Chen N, Winer DA (2023)."Chronic inflammation and the hallmarks of aging".Molecular Metabolism.74:101755.doi:10.1016/j.molmet.2023.101755.PMC10359950.PMID37329949.
- ^abLamming DW, Ye L, Katajisto P, et al. (March 2012)."Rapamycin-induced insulin resistance is mediated by mTORC2 loss and uncoupled from longevity".Science.335(6076): 1638–43.Bibcode:2012Sci...335.1638L.doi:10.1126/science.1215135.PMC3324089.PMID22461615.
- ^Zinzalla V, Stracka D, Oppliger W, et al. (March 2011)."Activation of mTORC2 by association with the ribosome".Cell.144(5): 757–68.doi:10.1016/j.cell.2011.02.014.PMID21376236.
- ^Zhang F, Zhang X, Li M, et al. (November 2010)."mTOR complex component Rictor interacts with PKCzeta and regulates cancer cell metastasis".Cancer Research.70(22): 9360–70.doi:10.1158/0008-5472.CAN-10-0207.PMID20978191.
- ^Guertin DA, Stevens DM, Thoreen CC, et al. (December 2006)."Ablation in mice of the mTORC components raptor, rictor, or mLST8 reveals that mTORC2 is required for signaling to Akt-FOXO and PKCalpha, but not S6K1".Developmental Cell.11(6): 859–71.doi:10.1016/j.devcel.2006.10.007.PMID17141160.
- ^Gu Y, Lindner J, Kumar A, et al. (March 2011)."Rictor/mTORC2 is essential for maintaining a balance between beta-cell proliferation and cell size".Diabetes.60(3): 827–37.doi:10.2337/db10-1194.PMC3046843.PMID21266327.
- ^Lamming DW, Demirkan G, Boylan JM, et al. (January 2014)."Hepatic signaling by the mechanistic target of rapamycin complex 2 (mTORC2)".FASEB Journal.28(1): 300–15.doi:10.1096/fj.13-237743.PMC3868844.PMID24072782.
- ^Kumar A, Lawrence JC, Jung DY, et al. (June 2010)."Fat cell-specific ablation of rictor in mice impairs insulin-regulated fat cell and whole-body glucose and lipid metabolism".Diabetes.59(6): 1397–406.doi:10.2337/db09-1061.PMC2874700.PMID20332342.
- ^Lamming DW, Mihaylova MM, Katajisto P, et al. (October 2014)."Depletion of Rictor, an essential protein component of mTORC2, decreases male lifespan".Aging Cell.13(5): 911–7.doi:10.1111/acel.12256.PMC4172536.PMID25059582.
- ^Feldman ME, Apsel B, Uotila A, et al. (February 2009)."Active-site inhibitors of mTOR target rapamycin-resistant outputs of mTORC1 and mTORC2".PLOS Biology.7(2): e38.doi:10.1371/journal.pbio.1000038.PMC2637922.PMID19209957.
- ^Wu JJ, Liu J, Chen EB, et al. (September 2013)."Increased mammalian lifespan and a segmental and tissue-specific slowing of aging after genetic reduction of mTOR expression".Cell Reports.4(5): 913–20.doi:10.1016/j.celrep.2013.07.030.PMC3784301.PMID23994476.
- ^Lawlor MA, Mora A, Ashby PR, et al. (July 2002)."Essential role of PDK1 in regulating cell size and development in mice".The EMBO Journal.21(14): 3728–38.doi:10.1093/emboj/cdf387.PMC126129.PMID12110585.
- ^Yang ZZ, Tschopp O, Baudry A, et al. (April 2004). "Physiological functions of protein kinase B/Akt".Biochemical Society Transactions.32(Pt 2): 350–4.doi:10.1042/BST0320350.PMID15046607.
- ^Nojima A, Yamashita M, Yoshida Y, et al. (2013-01-01)."Haploinsufficiency of akt1 prolongs the lifespan of mice".PLOS ONE.8(7): e69178.Bibcode:2013PLoSO...869178N.doi:10.1371/journal.pone.0069178.PMC3728301.PMID23935948.
- ^Crespo JL, Hall MN (December 2002)."Elucidating TOR signaling and rapamycin action: lessons from Saccharomyces cerevisiae".Microbiology and Molecular Biology Reviews.66(4): 579–91, table of contents.doi:10.1128/mmbr.66.4.579-591.2002.PMC134654.PMID12456783.
- ^Peter GJ, Düring L, Ahmed A (March 2006)."Carbon catabolite repression regulates amino acid permeases in Saccharomyces cerevisiae via the TOR signaling pathway".The Journal of Biological Chemistry.281(9): 5546–52.doi:10.1074/jbc.M513842200.PMID16407266.
- ^abPowers RW, Kaeberlein M, Caldwell SD, et al. (January 2006)."Extension of chronological life span in yeast by decreased TOR pathway signaling".Genes & Development.20(2): 174–84.doi:10.1101/gad.1381406.PMC1356109.PMID16418483.
- ^abKaeberlein M, Powers RW, Steffen KK, et al. (November 2005). "Regulation of yeast replicative life span by TOR and Sch9 in response to nutrients".Science.310(5751): 1193–6.Bibcode:2005Sci...310.1193K.doi:10.1126/science.1115535.PMID16293764.S2CID42188272.
- ^Jia K, Chen D, Riddle DL (August 2004). "The TOR pathway interacts with the insulin signaling pathway to regulate C. elegans larval development, metabolism and life span".Development.131(16): 3897–906.doi:10.1242/dev.01255.PMID15253933.S2CID10377667.
- ^Kapahi P, Zid BM, Harper T, et al. (May 2004)."Regulation of lifespan in Drosophila by modulation of genes in the TOR signaling pathway".Current Biology.14(10): 885–90.Bibcode:2004CBio...14..885K.doi:10.1016/j.cub.2004.03.059.PMC2754830.PMID15186745.
- ^Harrison DE, Strong R, Sharp ZD, et al. (July 2009)."Rapamycin fed late in life extends lifespan in genetically heterogeneous mice".Nature.460(7253): 392–5.Bibcode:2009Natur.460..392H.doi:10.1038/nature08221.PMC2786175.PMID19587680.
- ^Miller RA, Harrison DE, Astle CM, et al. (June 2014)."Rapamycin-mediated lifespan increase in mice is dose and sex dependent and metabolically distinct from dietary restriction".Aging Cell.13(3): 468–77.doi:10.1111/acel.12194.PMC4032600.PMID24341993.
- ^Fok WC, Chen Y, Bokov A, et al. (2014-01-01)."Mice fed rapamycin have an increase in lifespan associated with major changes in the liver transcriptome".PLOS ONE.9(1): e83988.Bibcode:2014PLoSO...983988F.doi:10.1371/journal.pone.0083988.PMC3883653.PMID24409289.
- ^Arriola Apelo SI, Pumper CP, Baar EL, et al. (July 2016)."Intermittent Administration of Rapamycin Extends the Life Span of Female C57BL/6J Mice".The Journals of Gerontology. Series A, Biological Sciences and Medical Sciences.71(7): 876–81.doi:10.1093/gerona/glw064.PMC4906329.PMID27091134.
- ^Popovich IG, Anisimov VN, Zabezhinski MA, et al. (May 2014)."Lifespan extension and cancer prevention in HER-2/neu transgenic mice treated with low intermittent doses of rapamycin".Cancer Biology & Therapy.15(5): 586–92.doi:10.4161/cbt.28164.PMC4026081.PMID24556924.
- ^Baar EL, Carbajal KA, Ong IM, et al. (February 2016)."Sex- and tissue-specific changes in mTOR signaling with age in C57BL/6J mice".Aging Cell.15(1): 155–66.doi:10.1111/acel.12425.PMC4717274.PMID26695882.
- ^Caron A, Richard D, Laplante M (Jul 2015). "The Roles of mTOR Complexes in Lipid Metabolism".Annual Review of Nutrition.35:321–48.doi:10.1146/annurev-nutr-071714-034355.PMID26185979.
- ^Cota D, Proulx K, Smith KA, et al. (May 2006). "Hypothalamic mTOR signaling regulates food intake".Science.312(5775): 927–30.Bibcode:2006Sci...312..927C.doi:10.1126/science.1124147.PMID16690869.S2CID6526786.
- ^abKriete A, Bosl WJ, Booker G (June 2010)."Rule-based cell systems model of aging using feedback loop motifs mediated by stress responses".PLOS Computational Biology.6(6): e1000820.Bibcode:2010PLSCB...6E0820K.doi:10.1371/journal.pcbi.1000820.PMC2887462.PMID20585546.
- ^abSchieke SM, Phillips D, McCoy JP, et al. (September 2006)."The mammalian target of rapamycin (mTOR) pathway regulates mitochondrial oxygen consumption and oxidative capacity".The Journal of Biological Chemistry.281(37): 27643–52.doi:10.1074/jbc.M603536200.PMID16847060.
- ^Yessenkyzy A, Saliev T, Zhanaliyeva M, et al. (2020)."Polyphenols as Caloric-Restriction Mimetics and Autophagy Inducers in Aging Research".Nutrients.12(5): 1344.doi:10.3390/nu12051344.PMC7285205.PMID32397145.
- ^abLaberge R, Sun Y, Orjalo AV, et al. (2015)."MTOR regulates the pro-tumorigenic senescence-associated secretory phenotype by promoting IL1A translation".Nature Cell Biology.17(8): 1049–1061.doi:10.1038/ncb3195.PMC4691706.PMID26147250.
- ^Wang R, Yu Z, Sunchu B, et al. (2017)."Rapamycin inhibits the secretory phenotype of senescent cells by a Nrf2-independent mechanism".Aging Cell.16(3): 564–574.doi:10.1111/acel.12587.PMC5418203.PMID28371119.
- ^Wang R, Sunchu B, Perez VI (2017). "Rapamycin and the inhibition of the secretory phenotype".Experimental Gerontology.94:89–92.doi:10.1016/j.exger.2017.01.026.PMID28167236.S2CID4960885.
- ^Weichhart T (2018)."mTOR as Regulator of Lifespan, Aging, and Cellular Senescence: A Mini-Review".Gerontology.84(2): 127–134.doi:10.1159/000484629.PMC6089343.PMID29190625.
- ^Xu K, Liu P, Wei W (December 2014)."mTOR signaling in tumorigenesis".Biochimica et Biophysica Acta (BBA) - Reviews on Cancer.1846(2): 638–54.doi:10.1016/j.bbcan.2014.10.007.PMC4261029.PMID25450580.
- ^Guertin DA, Sabatini DM (August 2005). "An expanding role for mTOR in cancer".Trends in Molecular Medicine.11(8): 353–61.doi:10.1016/j.molmed.2005.06.007.PMID16002336.
- ^Pópulo H, Lopes JM, Soares P (2012)."The mTOR signalling pathway in human cancer".International Journal of Molecular Sciences.13(2): 1886–918.doi:10.3390/ijms13021886.PMC3291999.PMID22408430.
- ^Easton JB, Houghton PJ (October 2006). "mTOR and cancer therapy".Oncogene.25(48): 6436–46.doi:10.1038/sj.onc.1209886.PMID17041628.S2CID19250234.
- ^Zoncu R, Efeyan A, Sabatini DM (January 2011)."mTOR: from growth signal integration to cancer, diabetes and ageing".Nature Reviews Molecular Cell Biology.12(1): 21–35.doi:10.1038/nrm3025.PMC3390257.PMID21157483.
- ^Thomas GV, Tran C, Mellinghoff IK, et al. (January 2006). "Hypoxia-inducible factor determines sensitivity to inhibitors of mTOR in kidney cancer".Nature Medicine.12(1): 122–7.doi:10.1038/nm1337.PMID16341243.S2CID1853822.
- ^Nemazanyy I, Espeillac C, Pende M, et al. (August 2013). "Role of PI3K, mTOR and Akt2 signalling in hepatic tumorigenesis via the control of PKM2 expression".Biochemical Society Transactions.41(4): 917–22.doi:10.1042/BST20130034.PMID23863156.
- ^Tang G, Gudsnuk K, Kuo SH, et al. (September 2014)."Loss of mTOR-dependent macroautophagy causes autistic-like synaptic pruning deficits".Neuron.83(5): 1131–43.doi:10.1016/j.neuron.2014.07.040.PMC4159743.PMID25155956.
- ^Rosner M, Hanneder M, Siegel N, et al. (June 2008). "The mTOR pathway and its role in human genetic diseases".Mutation Research.659(3): 284–92.doi:10.1016/j.mrrev.2008.06.001.PMID18598780.
- ^Li X, Alafuzoff I, Soininen H, et al. (August 2005). "Levels of mTOR and its downstream targets 4E-BP1, eEF2, and eEF2 kinase in relationships with tau in Alzheimer's disease brain".The FEBS Journal.272(16): 4211–20.doi:10.1111/j.1742-4658.2005.04833.x.PMID16098202.S2CID43085490.
- ^Chano T, Okabe H, Hulette CM (September 2007). "RB1CC1 insufficiency causes neuronal atrophy through mTOR signaling alteration and involved in the pathology of Alzheimer's diseases".Brain Research.1168(1168): 97–105.doi:10.1016/j.brainres.2007.06.075.PMID17706618.S2CID54255848.
- ^Selkoe DJ (September 2008)."Soluble oligomers of the amyloid beta-protein impair synaptic plasticity and behavior".Behavioural Brain Research.192(1): 106–13.doi:10.1016/j.bbr.2008.02.016.PMC2601528.PMID18359102.
- ^abOddo S (January 2012)."The role of mTOR signaling in Alzheimer disease".Frontiers in Bioscience.4(1): 941–52.doi:10.2741/s310.PMC4111148.PMID22202101.
- ^abAn WL, Cowburn RF, Li L, et al. (August 2003)."Up-regulation of phosphorylated/activated p70 S6 kinase and its relationship to neurofibrillary pathology in Alzheimer's disease".The American Journal of Pathology.163(2): 591–607.doi:10.1016/S0002-9440(10)63687-5.PMC1868198.PMID12875979.
- ^Zhang F, Beharry ZM, Harris TE, et al. (May 2009). "PIM1 protein kinase regulates PRAS40 phosphorylation and mTOR activity in FDCP1 cells".Cancer Biology & Therapy.8(9): 846–53.doi:10.4161/cbt.8.9.8210.PMID19276681.S2CID22153842.
- ^Koo EH, Squazzo SL (July 1994)."Evidence that production and release of amyloid beta-protein involves the endocytic pathway".The Journal of Biological Chemistry.269(26): 17386–9.doi:10.1016/S0021-9258(17)32449-3.PMID8021238.
- ^abcCaccamo A, Majumder S, Richardson A, et al. (April 2010)."Molecular interplay between mammalian target of rapamycin (mTOR), amyloid-beta, and Tau: effects on cognitive impairments".The Journal of Biological Chemistry.285(17): 13107–20.doi:10.1074/jbc.M110.100420.PMC2857107.PMID20178983.
- ^Lafay-Chebassier C, Paccalin M, Page G, et al. (July 2005)."mTOR/p70S6k signalling alteration by Abeta exposure as well as in APP-PS1 transgenic models and in patients with Alzheimer's disease".Journal of Neurochemistry.94(1): 215–25.doi:10.1111/j.1471-4159.2005.03187.x.PMID15953364.S2CID8464608.
- ^abcdCaccamo A, Maldonado MA, Majumder S, et al. (March 2011)."Naturally secreted amyloid-beta increases mammalian target of rapamycin (mTOR) activity via a PRAS40-mediated mechanism".The Journal of Biological Chemistry.286(11): 8924–32.doi:10.1074/jbc.M110.180638.PMC3058958.PMID21266573.
- ^Sancak Y, Thoreen CC, Peterson TR, et al. (March 2007)."PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase".Molecular Cell.25(6): 903–15.doi:10.1016/j.molcel.2007.03.003.PMID17386266.
- ^Wang L, Harris TE, Roth RA, et al. (July 2007)."PRAS40 regulates mTORC1 kinase activity by functioning as a direct inhibitor of substrate binding".The Journal of Biological Chemistry.282(27): 20036–44.doi:10.1074/jbc.M702376200.PMID17510057.
- ^Pei JJ, Hugon J (December 2008)."mTOR-dependent signalling in Alzheimer's disease".Journal of Cellular and Molecular Medicine.12(6B): 2525–32.doi:10.1111/j.1582-4934.2008.00509.x.PMC3828871.PMID19210753.
- ^Meske V, Albert F, Ohm TG (January 2008)."Coupling of mammalian target of rapamycin with phosphoinositide 3-kinase signaling pathway regulates protein phosphatase 2A- and glycogen synthase kinase-3 -dependent phosphorylation of Tau".The Journal of Biological Chemistry.283(1): 100–9.doi:10.1074/jbc.M704292200.PMID17971449.
- ^Janssens V, Goris J (February 2001)."Protein phosphatase 2A: a highly regulated family of serine/threonine phosphatases implicated in cell growth and signalling".The Biochemical Journal.353(Pt 3): 417–39.doi:10.1042/0264-6021:3530417.PMC1221586.PMID11171037.
- ^Morita T, Sobue K (October 2009)."Specification of neuronal polarity regulated by local translation of CRMP2 and Tau via the mTOR-p70S6K pathway".The Journal of Biological Chemistry.284(40): 27734–45.doi:10.1074/jbc.M109.008177.PMC2785701.PMID19648118.
- ^Puighermanal E, Marsicano G, Busquets-Garcia A, et al. (September 2009). "Cannabinoid modulation of hippocampal long-term memory is mediated by mTOR signaling".Nature Neuroscience.12(9): 1152–8.doi:10.1038/nn.2369.PMID19648913.S2CID9584832.
- ^Tischmeyer W, Schicknick H, Kraus M, et al. (August 2003). "Rapamycin-sensitive signalling in long-term consolidation of auditory cortex-dependent memory".The European Journal of Neuroscience.18(4): 942–50.doi:10.1046/j.1460-9568.2003.02820.x.PMID12925020.S2CID2780242.
- ^Hoeffer CA, Klann E (February 2010)."mTOR signaling: at the crossroads of plasticity, memory and disease".Trends in Neurosciences.33(2): 67–75.doi:10.1016/j.tins.2009.11.003.PMC2821969.PMID19963289.
- ^Kelleher RJ, Govindarajan A, Jung HY, et al. (February 2004)."Translational control by MAPK signaling in long-term synaptic plasticity and memory".Cell.116(3): 467–79.doi:10.1016/S0092-8674(04)00115-1.PMID15016380.
- ^Ehninger D, Han S, Shilyansky C, et al. (August 2008)."Reversal of learning deficits in a Tsc2+/- mouse model of tuberous sclerosis".Nature Medicine.14(8): 843–8.doi:10.1038/nm1788.PMC2664098.PMID18568033.
- ^Moreno JA, Radford H, Peretti D, et al. (May 2012)."Sustained translational repression by eIF2α-P mediates prion neurodegeneration".Nature.485(7399): 507–11.Bibcode:2012Natur.485..507M.doi:10.1038/nature11058.PMC3378208.PMID22622579.
- ^Díaz-Troya S, Pérez-Pérez ME, Florencio FJ, et al. (October 2008)."The role of TOR in autophagy regulation from yeast to plants and mammals".Autophagy.4(7): 851–65.doi:10.4161/auto.6555.PMID18670193.
- ^McCray BA, Taylor JP (December 2008). "The role of autophagy in age-related neurodegeneration".Neuro-Signals.16(1): 75–84.doi:10.1159/000109761.PMID18097162.S2CID13591350.
- ^Nedelsky NB, Todd PK, Taylor JP (December 2008)."Autophagy and the ubiquitin-proteasome system: collaborators in neuroprotection".Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease.1782(12): 691–9.doi:10.1016/j.bbadis.2008.10.002.PMC2621359.PMID18930136.
- ^Rubinsztein DC (October 2006). "The roles of intracellular protein-degradation pathways in neurodegeneration".Nature.443(7113): 780–6.Bibcode:2006Natur.443..780R.doi:10.1038/nature05291.PMID17051204.S2CID4411895.
- ^Oddo S (April 2008)."The ubiquitin-proteasome system in Alzheimer's disease".Journal of Cellular and Molecular Medicine.12(2): 363–73.doi:10.1111/j.1582-4934.2008.00276.x.PMC3822529.PMID18266959.
- ^Li X, Li H, Li XJ (November 2008)."Intracellular degradation of misfolded proteins in polyglutamine neurodegenerative diseases".Brain Research Reviews.59(1): 245–52.doi:10.1016/j.brainresrev.2008.08.003.PMC2577582.PMID18773920.
- ^Caccamo A, Majumder S, Deng JJ, et al. (October 2009)."Rapamycin rescues TDP-43 mislocalization and the associated low molecular mass neurofilament instability".The Journal of Biological Chemistry.284(40): 27416–24.doi:10.1074/jbc.M109.031278.PMC2785671.PMID19651785.
- ^Ravikumar B, Vacher C, Berger Z, et al. (June 2004)."Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease".Nature Genetics.36(6): 585–95.doi:10.1038/ng1362.PMID15146184.
- ^Rami A (October 2009)."Review: autophagy in neurodegeneration: firefighter and/or incendiarist?".Neuropathology and Applied Neurobiology.35(5): 449–61.doi:10.1111/j.1365-2990.2009.01034.x.PMID19555462.
- ^Völkl, Simon, et al. "Hyperactive mTOR pathway promotes lymphoproliferation and abnormal differentiation in autoimmune lymphoproliferative syndrome." Blood, The Journal of the American Society of Hematology 128.2 (2016): 227-238.https://doi.org/10.1182/blood-2015-11-685024
- ^Arenas, Daniel J., et al. "Increased mTOR activation in idiopathic multicentric Castleman disease." Blood 135.19 (2020): 1673-1684.https://doi.org/10.1182/blood.2019002792
- ^El-Salem, Mouna, et al. "Constitutive activation of mTOR signaling pathway in post-transplant lymphoproliferative disorders." Laboratory Investigation 87.1 (2007): 29-39.https://doi.org/10.1038/labinvest.3700494
- ^abcdBrook MS, Wilkinson DJ, Phillips BE, et al. (January 2016)."Skeletal muscle homeostasis and plasticity in youth and ageing: impact of nutrition and exercise".Acta Physiologica.216(1): 15–41.doi:10.1111/apha.12532.PMC4843955.PMID26010896.
- ^abBrioche T, Pagano AF, Py G, et al. (April 2016)."Muscle wasting and aging: Experimental models, fatty infiltrations, and prevention"(PDF).Molecular Aspects of Medicine.50:56–87.doi:10.1016/j.mam.2016.04.006.PMID27106402.S2CID29717535.
- ^Drummond MJ, Dreyer HC, Fry CS, et al. (April 2009)."Nutritional and contractile regulation of human skeletal muscle protein synthesis and mTORC1 signaling".Journal of Applied Physiology.106(4): 1374–84.doi:10.1152/japplphysiol.91397.2008.PMC2698645.PMID19150856.
- ^Salto R, Vílchez JD, Girón MD, et al. (2015)."β-Hydroxy-β-Methylbutyrate (HMB) Promotes Neurite Outgrowth in Neuro2a Cells".PLOS ONE.10(8): e0135614.Bibcode:2015PLoSO..1035614S.doi:10.1371/journal.pone.0135614.PMC4534402.PMID26267903.
- ^Kougias DG, Nolan SO, Koss WA, et al. (April 2016). "Beta-hydroxy-beta-methylbutyrate ameliorates aging effects in the dendritic tree of pyramidal neurons in the medial prefrontal cortex of both male and female rats".Neurobiology of Aging.40:78–85.doi:10.1016/j.neurobiolaging.2016.01.004.PMID26973106.S2CID3953100.
- ^abPhillips SM (May 2014)."A brief review of critical processes in exercise-induced muscular hypertrophy".Sports Med.44(Suppl 1): S71–S77.doi:10.1007/s40279-014-0152-3.PMC4008813.PMID24791918.
- ^abcdefgJia J, Abudu YP, Claude-Taupin A, et al. (April 2018)."Galectins Control mTOR in Response to Endomembrane Damage".Molecular Cell.70(1): 120–135.e8.doi:10.1016/j.molcel.2018.03.009.PMC5911935.PMID29625033.
- ^Noda T, Ohsumi Y (February 1998)."Tor, a phosphatidylinositol kinase homologue, controls autophagy in yeast".The Journal of Biological Chemistry.273(7): 3963–6.doi:10.1074/jbc.273.7.3963.PMID9461583.
- ^Dubouloz F, Deloche O, Wanke V, et al. (July 2005)."The TOR and EGO protein complexes orchestrate microautophagy in yeast".Molecular Cell.19(1): 15–26.doi:10.1016/j.molcel.2005.05.020.PMID15989961.
- ^abGanley IG, Lam du H, Wang J, et al. (May 2009)."ULK1.ATG13.FIP200 complex mediates mTOR signaling and is essential for autophagy".The Journal of Biological Chemistry.284(18): 12297–305.doi:10.1074/jbc.M900573200.PMC2673298.PMID19258318.
- ^abJung CH, Jun CB, Ro SH, et al. (April 2009)."ULK-Atg13-FIP200 complexes mediate mTOR signaling to the autophagy machinery".Molecular Biology of the Cell.20(7): 1992–2003.doi:10.1091/mbc.e08-12-1249.PMC2663920.PMID19225151.
- ^abHosokawa N, Hara T, Kaizuka T, et al. (April 2009)."Nutrient-dependent mTORC1 association with the ULK1-Atg13-FIP200 complex required for autophagy".Molecular Biology of the Cell.20(7): 1981–91.doi:10.1091/mbc.e08-12-1248.PMC2663915.PMID19211835.
- ^Hasegawa J, Maejima I, Iwamoto R, et al. (March 2015)."Selective autophagy: lysophagy".Methods.75:128–32.doi:10.1016/j.ymeth.2014.12.014.PMID25542097.
- ^Fraiberg M, Elazar Z (October 2016)."A TRIM16-Galactin3 Complex Mediates Autophagy of Damaged Endomembranes".Developmental Cell.39(1): 1–2.doi:10.1016/j.devcel.2016.09.025.PMID27728777.
- ^abcChauhan S, Kumar S, Jain A, et al. (October 2016)."TRIMs and Galectins Globally Cooperate and TRIM16 and Galectin-3 Co-direct Autophagy in Endomembrane Damage Homeostasis".Developmental Cell.39(1): 13–27.doi:10.1016/j.devcel.2016.08.003.PMC5104201.PMID27693506.
- ^Nishimura T, Kaizuka T, Cadwell K, et al. (March 2013)."FIP200 regulates targeting of Atg16L1 to the isolation membrane".EMBO Reports.14(3): 284–91.doi:10.1038/embor.2013.6.PMC3589088.PMID23392225.
- ^Gammoh N, Florey O, Overholtzer M, et al. (February 2013)."Interaction between FIP200 and ATG16L1 distinguishes ULK1 complex-dependent and -independent autophagy".Nature Structural & Molecular Biology.20(2): 144–9.doi:10.1038/nsmb.2475.PMC3565010.PMID23262492.
- ^abcFujita N, Morita E, Itoh T, et al. (October 2013)."Recruitment of the autophagic machinery to endosomes during infection is mediated by ubiquitin".The Journal of Cell Biology.203(1): 115–28.doi:10.1083/jcb.201304188.PMC3798248.PMID24100292.
- ^Park JM, Jung CH, Seo M, et al. (2016-03-03)."The ULK1 complex mediates MTORC1 signaling to the autophagy initiation machinery via binding and phosphorylating ATG14".Autophagy.12(3): 547–64.doi:10.1080/15548627.2016.1140293.PMC4835982.PMID27046250.
- ^Dooley HC, Razi M, Polson HE, et al. (July 2014)."WIPI2 links LC3 conjugation with PI3P, autophagosome formation, and pathogen clearance by recruiting Atg12-5-16L1".Molecular Cell.55(2): 238–52.doi:10.1016/j.molcel.2014.05.021.PMC4104028.PMID24954904.
- ^Kim J, Kundu M, Viollet B, et al. (February 2011)."AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1".Nature Cell Biology.13(2): 132–41.doi:10.1038/ncb2152.PMC3987946.PMID21258367.
- ^Kim J, Kim YC, Fang C, et al. (January 2013)."Differential regulation of distinct Vps34 complexes by AMPK in nutrient stress and autophagy".Cell.152(1–2): 290–303.doi:10.1016/j.cell.2012.12.016.PMC3587159.PMID23332761.
- ^Gwinn DM, Shackelford DB, Egan DF, et al. (April 2008)."AMPK phosphorylation of raptor mediates a metabolic checkpoint".Molecular Cell.30(2): 214–26.doi:10.1016/j.molcel.2008.03.003.PMC2674027.PMID18439900.
- ^Inoki K, Zhu T, Guan KL (November 2003)."TSC2 mediates cellular energy response to control cell growth and survival".Cell.115(5): 577–90.doi:10.1016/S0092-8674(03)00929-2.PMID14651849.
- ^Yoshida Y, Yasuda S, Fujita T, et al. (August 2017)."FBXO27 directs damaged lysosomes for autophagy".Proceedings of the National Academy of Sciences of the United States of America.114(32): 8574–8579.doi:10.1073/pnas.1702615114.PMC5559013.PMID28743755.
- ^Jimenez SA, Cronin PM, Koenig AS, et al. (15 February 2012). Varga J, Talavera F, Goldberg E, Mechaber AJ, Diamond HS (eds.)."Scleroderma".Medscape Reference.WebMD.Retrieved5 March2014.
- ^Hajj-ali RA (June 2013)."Systemic Sclerosis".Merck Manual Professional.Merck Sharp & Dohme Corp.Retrieved5 March2014.
- ^"Mammalian target of rapamycin (mTOR) inhibitors in solid tumours".Pharmaceutical Journal.Retrieved2018-10-18.
- ^Faivre S, Kroemer G, Raymond E (August 2006). "Current development of mTOR inhibitors as anticancer agents".Nature Reviews. Drug Discovery.5(8): 671–88.doi:10.1038/nrd2062.PMID16883305.S2CID27952376.
- ^Hasty P (February 2010)."Rapamycin: the cure for all that ails".Journal of Molecular Cell Biology.2(1): 17–9.doi:10.1093/jmcb/mjp033.PMID19805415.
- ^Bové J, Martínez-Vicente M, Vila M (August 2011). "Fighting neurodegeneration with rapamycin: mechanistic insights".Nature Reviews. Neuroscience.12(8): 437–52.doi:10.1038/nrn3068.PMID21772323.S2CID205506774.
- ^Mannick JB, Morris M, Hockey HP, et al. (July 2018)."TORC1 inhibition enhances immune function and reduces infections in the elderly".Science Translational Medicine.10(449): eaaq1564.doi:10.1126/scitranslmed.aaq1564.PMID29997249.
- ^Estrela JM, Ortega A, Mena S, et al. (2013). "Pterostilbene: Biomedical applications".Critical Reviews in Clinical Laboratory Sciences.50(3): 65–78.doi:10.3109/10408363.2013.805182.PMID23808710.S2CID45618964.
- ^McCubrey JA, Lertpiriyapong K, Steelman LS, et al. (June 2017)."Effects of resveratrol, curcumin, berberine and other nutraceuticals on aging, cancer development, cancer stem cells and microRNAs".Aging.9(6): 1477–1536.doi:10.18632/aging.101250.PMC5509453.PMID28611316.
- ^Malavolta M, Bracci M, Santarelli L, et al. (2018)."Inducers of Senescence, Toxic Compounds, and Senolytics: The Multiple Faces of Nrf2-Activating Phytochemicals in Cancer Adjuvant Therapy".Mediators of Inflammation.2018:4159013.doi:10.1155/2018/4159013.PMC5829354.PMID29618945.
- ^Gómez-Linton DR, Alavez S, Alarcón-Aguilar A, et al. (October 2019). "Some naturally occurring compounds that increase longevity and stress resistance in model organisms of aging".Biogerontology.20(5): 583–603.doi:10.1007/s10522-019-09817-2.PMID31187283.S2CID184483900.
- ^Li W, Qin L, Feng R, et al. (July 2019). "Emerging senolytic agents derived from natural products".Mechanisms of Ageing and Development.181:1–6.doi:10.1016/j.mad.2019.05.001.PMID31077707.S2CID147704626.
- ^"mTOR protein interactors".Human Protein Reference Database.Johns Hopkins University and the Institute of Bioinformatics. Archived fromthe originalon 2015-06-28.Retrieved2010-12-06.
- ^Kumar V, Sabatini D, Pandey P, et al. (April 2000)."Regulation of the rapamycin and FKBP-target 1/mammalian target of rapamycin and cap-dependent initiation of translation by the c-Abl protein-tyrosine kinase".The Journal of Biological Chemistry.275(15): 10779–87.doi:10.1074/jbc.275.15.10779.PMID10753870.
- ^Sekulić A, Hudson CC, Homme JL, et al. (July 2000). "A direct linkage between the phosphoinositide 3-kinase-AKT signaling pathway and the mammalian target of rapamycin in mitogen-stimulated and transformed cells".Cancer Research.60(13): 3504–13.PMID10910062.
- ^Cheng SW, Fryer LG, Carling D, et al. (April 2004)."Thr2446 is a novel mammalian target of rapamycin (mTOR) phosphorylation site regulated by nutrient status".The Journal of Biological Chemistry.279(16): 15719–22.doi:10.1074/jbc.C300534200.PMID14970221.
- ^Choi JH, Bertram PG, Drenan R, et al. (October 2002)."The FKBP12-rapamycin-associated protein (FRAP) is a CLIP-170 kinase".EMBO Reports.3(10): 988–94.doi:10.1093/embo-reports/kvf197.PMC1307618.PMID12231510.
- ^Harris TE, Chi A, Shabanowitz J, et al. (April 2006)."mTOR-dependent stimulation of the association of eIF4G and eIF3 by insulin".The EMBO Journal.25(8): 1659–68.doi:10.1038/sj.emboj.7601047.PMC1440840.PMID16541103.
- ^abSchalm SS, Fingar DC, Sabatini DM, et al. (May 2003)."TOS motif-mediated raptor binding regulates 4E-BP1 multisite phosphorylation and function".Current Biology.13(10): 797–806.Bibcode:2003CBio...13..797S.doi:10.1016/S0960-9822(03)00329-4.PMID12747827.
- ^abcHara K, Maruki Y, Long X, et al. (July 2002)."Raptor, a binding partner of target of rapamycin (TOR), mediates TOR action".Cell.110(2): 177–89.doi:10.1016/S0092-8674(02)00833-4.PMID12150926.
- ^abWang L, Rhodes CJ, Lawrence JC (August 2006)."Activation of mammalian target of rapamycin (mTOR) by insulin is associated with stimulation of 4EBP1 binding to dimeric mTOR complex 1".The Journal of Biological Chemistry.281(34): 24293–303.doi:10.1074/jbc.M603566200.PMID16798736.
- ^abcLong X, Lin Y, Ortiz-Vega S, et al. (April 2005)."Rheb binds and regulates the mTOR kinase".Current Biology.15(8): 702–13.Bibcode:2005CBio...15..702L.doi:10.1016/j.cub.2005.02.053.PMID15854902.
- ^abTakahashi T, Hara K, Inoue H, et al. (September 2000). "Carboxyl-terminal region conserved among phosphoinositide-kinase-related kinases is indispensable for mTOR function in vivo and in vitro".Genes to Cells.5(9): 765–75.doi:10.1046/j.1365-2443.2000.00365.x.PMID10971657.S2CID39048740.
- ^abBurnett PE, Barrow RK, Cohen NA, et al. (February 1998)."RAFT1 phosphorylation of the translational regulators p70 S6 kinase and 4E-BP1".Proceedings of the National Academy of Sciences of the United States of America.95(4): 1432–7.Bibcode:1998PNAS...95.1432B.doi:10.1073/pnas.95.4.1432.PMC19032.PMID9465032.
- ^Wang X, Beugnet A, Murakami M, et al. (April 2005)."Distinct signaling events downstream of mTOR cooperate to mediate the effects of amino acids and insulin on initiation factor 4E-binding proteins".Molecular and Cellular Biology.25(7): 2558–72.doi:10.1128/MCB.25.7.2558-2572.2005.PMC1061630.PMID15767663.
- ^Choi J, Chen J, Schreiber SL, et al. (July 1996). "Structure of the FKBP12-rapamycin complex interacting with the binding domain of human FRAP".Science.273(5272): 239–42.Bibcode:1996Sci...273..239C.doi:10.1126/science.273.5272.239.PMID8662507.S2CID27706675.
- ^Luker KE, Smith MC, Luker GD, et al. (August 2004)."Kinetics of regulated protein-protein interactions revealed with firefly luciferase complementation imaging in cells and living animals".Proceedings of the National Academy of Sciences of the United States of America.101(33): 12288–93.Bibcode:2004PNAS..10112288L.doi:10.1073/pnas.0404041101.PMC514471.PMID15284440.
- ^Banaszynski LA, Liu CW, Wandless TJ (April 2005). "Characterization of the FKBP.rapamycin.FRB ternary complex".Journal of the American Chemical Society.127(13): 4715–21.doi:10.1021/ja043277y.PMID15796538.
- ^Sabers CJ, Martin MM, Brunn GJ, et al. (January 1995)."Isolation of a protein target of the FKBP12-rapamycin complex in mammalian cells".The Journal of Biological Chemistry.270(2): 815–22.doi:10.1074/jbc.270.2.815.PMID7822316.
- ^Sabatini DM, Barrow RK, Blackshaw S, et al. (May 1999). "Interaction of RAFT1 with gephyrin required for rapamycin-sensitive signaling".Science.284(5417): 1161–4.Bibcode:1999Sci...284.1161S.doi:10.1126/science.284.5417.1161.PMID10325225.
- ^Ha SH, Kim DH, Kim IS, et al. (December 2006). "PLD2 forms a functional complex with mTOR/raptor to transduce mitogenic signals".Cellular Signalling.18(12): 2283–91.doi:10.1016/j.cellsig.2006.05.021.PMID16837165.
- ^Buerger C, DeVries B, Stambolic V (June 2006). "Localization of Rheb to the endomembrane is critical for its signaling function".Biochemical and Biophysical Research Communications.344(3): 869–80.doi:10.1016/j.bbrc.2006.03.220.PMID16631613.
- ^abJacinto E, Facchinetti V, Liu D, et al. (October 2006)."SIN1/MIP1 maintains rictor-mTOR complex integrity and regulates Akt phosphorylation and substrate specificity".Cell.127(1): 125–37.doi:10.1016/j.cell.2006.08.033.PMID16962653.
- ^McMahon LP, Yue W, Santen RJ, et al. (January 2005)."Farnesylthiosalicylic acid inhibits mammalian target of rapamycin (mTOR) activity both in cells and in vitro by promoting dissociation of the mTOR-raptor complex".Molecular Endocrinology.19(1): 175–83.doi:10.1210/me.2004-0305.PMID15459249.
- ^Oshiro N, Yoshino K, Hidayat S, et al. (April 2004). "Dissociation of raptor from mTOR is a mechanism of rapamycin-induced inhibition of mTOR function".Genes to Cells.9(4): 359–66.doi:10.1111/j.1356-9597.2004.00727.x.hdl:20.500.14094/D1002969.PMID15066126.S2CID24814691.
- ^Kawai S, Enzan H, Hayashi Y, et al. (July 2003). "Vinculin: a novel marker for quiescent and activated hepatic stellate cells in human and rat livers".Virchows Archiv.443(1): 78–86.doi:10.1007/s00428-003-0804-4.PMID12719976.S2CID21552704.
- ^Choi KM, McMahon LP, Lawrence JC (May 2003)."Two motifs in the translational repressor PHAS-I required for efficient phosphorylation by mammalian target of rapamycin and for recognition by raptor".The Journal of Biological Chemistry.278(22): 19667–73.doi:10.1074/jbc.M301142200.PMID12665511.
- ^abNojima H, Tokunaga C, Eguchi S, et al. (May 2003)."The mammalian target of rapamycin (mTOR) partner, raptor, binds the mTOR substrates p70 S6 kinase and 4E-BP1 through their TOR signaling (TOS) motif".The Journal of Biological Chemistry.278(18): 15461–4.doi:10.1074/jbc.C200665200.PMID12604610.
- ^abSarbassov DD, Ali SM, Sengupta S, et al. (April 2006)."Prolonged rapamycin treatment inhibits mTORC2 assembly and Akt/PKB".Molecular Cell.22(2): 159–68.doi:10.1016/j.molcel.2006.03.029.PMID16603397.
- ^Tzatsos A, Kandror KV (January 2006)."Nutrients suppress phosphatidylinositol 3-kinase/Akt signaling via raptor-dependent mTOR-mediated insulin receptor substrate 1 phosphorylation".Molecular and Cellular Biology.26(1): 63–76.doi:10.1128/MCB.26.1.63-76.2006.PMC1317643.PMID16354680.
- ^abcSarbassov DD, Sabatini DM (November 2005)."Redox regulation of the nutrient-sensitive raptor-mTOR pathway and complex".The Journal of Biological Chemistry.280(47): 39505–9.doi:10.1074/jbc.M506096200.PMID16183647.
- ^abYang Q, Inoki K, Ikenoue T, et al. (October 2006)."Identification of Sin1 as an essential TORC2 component required for complex formation and kinase activity".Genes & Development.20(20): 2820–32.doi:10.1101/gad.1461206.PMC1619946.PMID17043309.
- ^Kumar V, Pandey P, Sabatini D, et al. (March 2000)."Functional interaction between RAFT1/FRAP/mTOR and protein kinase cdelta in the regulation of cap-dependent initiation of translation".The EMBO Journal.19(5): 1087–97.doi:10.1093/emboj/19.5.1087.PMC305647.PMID10698949.
- ^Long X, Ortiz-Vega S, Lin Y, et al. (June 2005)."Rheb binding to mammalian target of rapamycin (mTOR) is regulated by amino acid sufficiency".The Journal of Biological Chemistry.280(25): 23433–6.doi:10.1074/jbc.C500169200.PMID15878852.
- ^Smith EM, Finn SG, Tee AR, et al. (May 2005)."The tuberous sclerosis protein TSC2 is not required for the regulation of the mammalian target of rapamycin by amino acids and certain cellular stresses".The Journal of Biological Chemistry.280(19): 18717–27.doi:10.1074/jbc.M414499200.PMID15772076.
- ^Bernardi R, Guernah I, Jin D, et al. (August 2006). "PML inhibits HIF-1alpha translation and neoangiogenesis through repression of mTOR".Nature.442(7104): 779–85.Bibcode:2006Natur.442..779B.doi:10.1038/nature05029.PMID16915281.S2CID4427427.
- ^Saitoh M, Pullen N, Brennan P, et al. (May 2002)."Regulation of an activated S6 kinase 1 variant reveals a novel mammalian target of rapamycin phosphorylation site".The Journal of Biological Chemistry.277(22): 20104–12.doi:10.1074/jbc.M201745200.PMID11914378.
- ^Chiang GG, Abraham RT (July 2005)."Phosphorylation of mammalian target of rapamycin (mTOR) at Ser-2448 is mediated by p70S6 kinase".The Journal of Biological Chemistry.280(27): 25485–90.doi:10.1074/jbc.M501707200.PMID15899889.
- ^Holz MK, Blenis J (July 2005)."Identification of S6 kinase 1 as a novel mammalian target of rapamycin (mTOR)-phosphorylating kinase".The Journal of Biological Chemistry.280(28): 26089–93.doi:10.1074/jbc.M504045200.PMID15905173.
- ^Isotani S, Hara K, Tokunaga C, et al. (November 1999)."Immunopurified mammalian target of rapamycin phosphorylates and activates p70 S6 kinase alpha in vitro".The Journal of Biological Chemistry.274(48): 34493–8.doi:10.1074/jbc.274.48.34493.hdl:20.500.14094/D1002182.PMID10567431.
- ^Toral-Barza L, Zhang WG, Lamison C, et al. (June 2005). "Characterization of the cloned full-length and a truncated human target of rapamycin: activity, specificity, and enzyme inhibition as studied by a high capacity assay".Biochemical and Biophysical Research Communications.332(1): 304–10.doi:10.1016/j.bbrc.2005.04.117.PMID15896331.
- ^Ali SM, Sabatini DM (May 2005)."Structure of S6 kinase 1 determines whether raptor-mTOR or rictor-mTOR phosphorylates its hydrophobic motif site".The Journal of Biological Chemistry.280(20): 19445–8.doi:10.1074/jbc.C500125200.PMID15809305.
- ^Edinger AL, Linardic CM, Chiang GG, et al. (December 2003). "Differential effects of rapamycin on mammalian target of rapamycin signaling functions in mammalian cells".Cancer Research.63(23): 8451–60.PMID14679009.
- ^Leone M, Crowell KJ, Chen J, et al. (August 2006). "The FRB domain of mTOR: NMR solution structure and inhibitor design".Biochemistry.45(34): 10294–302.doi:10.1021/bi060976+.PMID16922504.
- ^Kristof AS, Marks-Konczalik J, Billings E, et al. (September 2003)."Stimulation of signal transducer and activator of transcription-1 (STAT1)-dependent gene transcription by lipopolysaccharide and interferon-gamma is regulated by mammalian target of rapamycin".The Journal of Biological Chemistry.278(36): 33637–44.doi:10.1074/jbc.M301053200.PMID12807916.
- ^Yokogami K, Wakisaka S, Avruch J, et al. (January 2000)."Serine phosphorylation and maximal activation of STAT3 during CNTF signaling is mediated by the rapamycin target mTOR".Current Biology.10(1): 47–50.Bibcode:2000CBio...10...47Y.doi:10.1016/S0960-9822(99)00268-7.PMID10660304.
- ^Kusaba H, Ghosh P, Derin R, et al. (January 2005)."Interleukin-12-induced interferon-gamma production by human peripheral blood T cells is regulated by mammalian target of rapamycin (mTOR)".The Journal of Biological Chemistry.280(2): 1037–43.doi:10.1074/jbc.M405204200.PMID15522880.
- ^Cang C, Zhou Y, Navarro B, et al. (February 2013)."mTOR regulates lysosomal ATP-sensitive two-pore Na(+) channels to adapt to metabolic state".Cell.152(4): 778–90.doi:10.1016/j.cell.2013.01.023.PMC3908667.PMID23394946.
- ^Wu S, Mikhailov A, Kallo-Hosein H, et al. (January 2002)."Characterization of ubiquilin 1, an mTOR-interacting protein".Biochimica et Biophysica Acta (BBA) - Molecular Cell Research.1542(1–3): 41–56.doi:10.1016/S0167-4889(01)00164-1.PMID11853878.
Further reading[edit]
- Saxton RA, Sabatini DM (March 2017)."mTOR Signaling in Growth, Metabolism, and Disease".Cell.168(6): 960–976.doi:10.1016/j.cell.2017.02.004.PMC5394987.PMID28283069.
External links[edit]
- mTOR+proteinat the U.S. National Library of MedicineMedical Subject Headings(MeSH)
- "mTOR Signaling Pathway in Pathway Interaction Database".National Cancer Institute. Archived fromthe originalon 2013-03-18.Retrieved2015-10-18.
- Overview of all the structural information available in thePDBforUniProt:P42345(Serine/threonine-protein kinase mTOR) at thePDBe-KB.