Steroid sulfatase
| STS | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | STS,ARSC, ARSC2, ARSC1, ASC, ES, SSDD, XLI, Steroid sulfatase (microsomal), isozyme S, steroid sulfatase | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM:300747;HomoloGene:47918;GeneCards:STS;OMA:STS - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Steryl-sulfatase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 3.1.6.2 | ||||||||
| CAS no. | 9025-62-1 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDBstructures | RCSB PDBPDBePDBsum | ||||||||
| |||||||||
Steroid sulfatase(STS), orsteryl-sulfatase(EC 3.1.6.2), formerly known asarylsulfatase C,is asulfataseenzymeinvolved in the metabolism ofsteroids.It is encoded by theSTSgene.[3]
Reactions[edit]
This enzymecatalysesthe followingchemical reaction
- 3β-hydroxyandrost-5-en-17-one 3-sulfate + H2O3β-hydroxyandrost-5-en-17-one +sulfate
Also acts on some related steryl sulfates.
Function[edit]
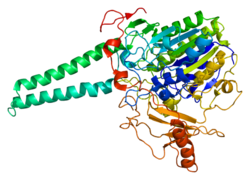
The protein encoded by this gene catalyzes the conversion ofsulfated steroidprecursors to the free steroid. This includesDHEA sulfate,estrone sulfate,pregnenolone sulfate,andcholesterol sulfate,all to their unconjugated forms (DHEA,estrone,pregnenolone,andcholesterol,respectively).[4][5]The encoded protein is found in theendoplasmic reticulum,where it is present as ahomodimer.[3]

Clinical significance[edit]
A congenital deficiency in the enzyme is associated withX-linked ichthyosis,a scaly-skin disease affecting roughly 1 in every 2,000 to 6,000 males.[7][8]The excessiveskin scalingandhyperkeratosisis caused by a lack of breakdown and thus accumulation of cholesterol sulfate, a steroid that stabilizes cell membranes and adds cohesion, in the outer layers of the skin.[4]
Genetic deletions including STS are associated with an increased risk of developmental and mood disorders (and associated traits), and ofatrial fibrillationoratrial flutterin males.[9]Both steroid sulfatase deficiency and common genetic risk variants within STS may confer increasedatrial fibrillationrisk.[10]Cardiacarrhythmiain STS deficiency may be related to abnormal development of theinterventricular septumorinteratrial septum.[11]Blood-clotting abnormalities may occur more frequently in males with XLI and female carriers.[12]Knockdown of STS gene expression in human skin cell cultures affects pathways associated with skin function, brain and heart development, and blood-clotting that may be relevant for explaining the skin condition and increased likelihood of ADHD/autism, cardiac arrhythmias and disorders ofhemostasisin XLI.[13]
Steroid sulfates like DHEA sulfate and estrone sulfate serve as large biologically inert reservoirs for conversion into androgens and estrogens, respectively, and hence are of significance forandrogen-andestrogen-dependent conditionslikeprostate cancer,breast cancer,endometriosis,and others. A number of clinical trials have been performed with inhibitors of the enzyme that have demonstrated clinical benefit, particularly in oncology and so far up to Phase II.[14]The non-steroidal drugIrosustathas been the most studied to date.
Inhibitors[edit]
Inhibitorsof STS includeirosustat,estrone sulfamate(EMATE),estradiol sulfamate(E2MATE), anddanazol.[15][16]The most potent inhibitors are based around the aryl sulfamate pharmacophore[17]and it is thought that such compounds irreversibly modify the active site formylglycine residue of steroid sulfatase.[14]
Names[edit]
Steryl-sulfatase is also known asarylsulfatase,steroid sulfatase,sterol sulfatase,dehydroepiandrosterone sulfate sulfatase,arylsulfatase C,steroid 3-sulfatase,steroid sulfate sulfohydrolase,dehydroepiandrosterone sulfatase,pregnenolone sulfatase,phenolic steroid sulfatase,3-beta-hydroxysteroid sulfate sulfatase,as well as by itssystematic namesteryl-sulfate sulfohydrolase.[18][19][20]
See also[edit]
References[edit]
- ^abcGRCh38: Ensembl release 89: ENSG00000101846–Ensembl,May 2017
- ^"Human PubMed Reference:".National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ab"Entrez Gene: STS steroid sulfatase (microsomal), arylsulfatase C, isozyme S".
- ^abMueller JW, Gilligan LC, Idkowiak J, Arlt W, Foster PA (October 2015)."The Regulation of Steroid Action by Sulfation and Desulfation".Endocrine Reviews.36(5): 526–63.doi:10.1210/er.2015-1036.PMC4591525.PMID26213785.
- ^Rižner TL (2016)."The Important Roles of Steroid Sulfatase and Sulfotransferases in Gynecological Diseases".Frontiers in Pharmacology.7:30.doi:10.3389/fphar.2016.00030.PMC4757672.PMID26924986.
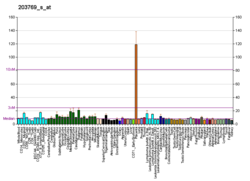
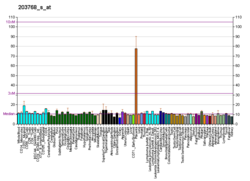
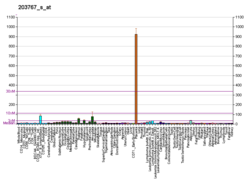
- ^Miki Y, Nakata T, Suzuki T, Darnel AD, Moriya T, Kaneko C, et al. (December 2002)."Systemic distribution of steroid sulfatase and estrogen sulfotransferase in human adult and fetal tissues".The Journal of Clinical Endocrinology and Metabolism.87(12): 5760–8.doi:10.1210/jc.2002-020670.PMID12466383.
- ^Alperin ES, Shapiro LJ (August 1997)."Characterization of point mutations in patients with X-linked ichthyosis. Effects on the structure and function of the steroid sulfatase protein".The Journal of Biological Chemistry.272(33): 20756–63.doi:10.1074/jbc.272.33.20756.PMID9252398.
- ^Ghosh D (December 2004). "Mutations in X-linked ichthyosis disrupt the active site structure of estrone/DHEA sulfatase".Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease.1739(1): 1–4.doi:10.1016/j.bbadis.2004.09.003.PMID15607112.
- ^Brcic L, Underwood JF, Kendall KM, Caseras X, Kirov G, Davies W (Mar 2020)."Medical and neurobehavioural phenotypes in carriers of X-linked ichthyosis-associated genetic deletions in the UK Biobank".Journal of Medical Genetics.57(10): 692–698.doi:10.1136/jmedgenet-2019-106676.PMC7525778.PMID32139392.
- ^Wren G, Baker E, Underwood J, Humby T, Thompson A, Kirov G, Escott-Price V, Davies W (November 2022)."Characterising heart rhythm abnormalities associated with Xp22.31 deletion".Journal of Medical Genetics.60(7): 636–643.doi:10.1136/jmg-2022-108862.PMC10359567.PMID36379544.
- ^Wren GH, Davies W (April 2024)."Cardiac arrhythmia in individuals with steroid sulfatase deficiency (X-linked ichthyosis): candidate anatomical and biochemical pathways".Essays in Biochemistry.doi:10.1042/EBC20230098.PMID38571328.
- ^Brcic L, Wren GH, Underwood JF, Kirov G, Davies W (May 2022)."Comorbid Medical Issues in X-Linked Ichthyosis".JID Innovations: Skin Science from Molecules to Population Health.2(3): 100109.doi:10.1016/j.xjidi.2022.100109.PMC8938907.PMID35330591.
- ^McGeoghan F, Camera E, Maiellaro M, Menon M, Huang M, Dewan P, Ziaj S, Caley MP, Donaldson M, Enright AJ, O'Toole EA (2023)."RNA sequencing and lipidomics uncovers novel pathomechanisms in recessive X-linked ichthyosis".Frontiers in Molecular Biosciences.10:1176802.doi:10.3389/fmolb.2023.1176802.PMC10285781.PMID37363400.
- ^abPotter BV (August 2018)."SULFATION PATHWAYS: Steroid sulphatase inhibition via aryl sulphamates: clinical progress, mechanism and future prospects".Journal of Molecular Endocrinology.61(2): T233–T252.doi:10.1530/JME-18-0045.PMID29618488.
- ^Thomas MP, Potter BV (September 2015). "EstrogenO-sulfamates and their analogues: Clinical steroid sulfatase inhibitors with broad potential ".The Journal of Steroid Biochemistry and Molecular Biology.153:160–9.doi:10.1016/j.jsbmb.2015.03.012.PMID25843211.S2CID24116740.
- ^Carlström K, Döberl A, Pousette A, Rannevik G, Wilking N (1984). "Inhibition of steroid sulfatase activity by danazol".Acta Obstetricia et Gynecologica Scandinavica Supplement.123:107–11.doi:10.3109/00016348409156994.PMID6238495.S2CID45817485.
- ^Thomas MP, Potter BV (October 2015)."Discovery and Development of the ArylO-Sulfamate Pharmacophore for Oncology and Women's Health ".Journal of Medicinal Chemistry.58(19): 7634–58.doi:10.1021/acs.jmedchem.5b00386.PMC5159624.PMID25992880.
- ^Roy AB (October 1954)."The steroid sulphatase ofPatella vulgata".Biochimica et Biophysica Acta.15(2): 300–1.doi:10.1016/0006-3002(54)90078-5.PMC1274509.PMID13208702.
- ^Roy AB (1960). "The Synthesis and Hydrolysis of Sulfate Esters".Advances in Enzymology and Related Areas of Molecular Biology.Advances in Enzymology - and Related Areas of Molecular Biology. Vol. 22. pp. 205–35.doi:10.1002/9780470122679.ch5.ISBN9780470122679.PMID13744184.
{{cite book}}:|journal=ignored (help) - ^Halkerston ID, Hillman J, Stitch SR (August 1956)."The enzymic hydrolysis of steroid conjugates. I. Sulphatase and β-glucuronidase activity of molluscan extracts".The Biochemical Journal.63(4): 705–10.doi:10.1042/bj0630705.PMC1216242.PMID13355874.
Further reading[edit]
- Elias PM, Crumrine D, Rassner U, Hachem JP, Menon GK, Man W, et al. (February 2004)."Basis for abnormal desquamation and permeability barrier dysfunction in RXLI".The Journal of Investigative Dermatology.122(2): 314–9.doi:10.1046/j.1523-1747.2003.22258.x.PMID15009711.
- Basler E, Grompe M, Parenti G, Yates J, Ballabio A (March 1992)."Identification of point mutations in the steroid sulfatase gene of three patients with X-linked ichthyosis".American Journal of Human Genetics.50(3): 483–91.PMC1684279.PMID1539590.
- Shankaran R, Ameen M, Daniel WL, Davidson RG, Chang PL (June 1991). "Characterization of arylsulfatase C isozymes from human liver and placenta".Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology.1078(2): 251–7.doi:10.1016/0167-4838(91)90566-I.PMID2065092.
- Stein C, Hille A, Seidel J, Rijnbout S, Waheed A, Schmidt B, et al. (August 1989)."Cloning and expression of human steroid-sulfatase. Membrane topology, glycosylation, and subcellular distribution in BHK-21 cells".The Journal of Biological Chemistry.264(23): 13865–72.doi:10.1016/S0021-9258(18)80080-1.PMID2668275.
- Kawano J, Kotani T, Ohtaki S, Minamino N, Matsuo H, Oinuma T, Aikawa E (August 1989). "Characterization of rat and human steroid sulfatases".Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology.997(3): 199–205.doi:10.1016/0167-4838(89)90187-8.PMID2765556.
- Yen PH, Allen E, Marsh B, Mohandas T, Wang N, Taggart RT, Shapiro LJ (May 1987). "Cloning and expression of steroid sulfatase cDNA and the frequent occurrence of deletions in STS deficiency: implications for X-Y interchange".Cell.49(4): 443–54.doi:10.1016/0092-8674(87)90447-8.PMID3032454.S2CID23353934.
- Conary JT, Lorkowski G, Schmidt B, Pohlmann R, Nagel G, Meyer HE, et al. (April 1987). "Genetic heterogeneity of steroid sulfatase deficiency revealed with cDNA for human steroid sulfatase".Biochemical and Biophysical Research Communications.144(2): 1010–7.doi:10.1016/S0006-291X(87)80064-5.PMID3034252.
- Ballabio A, Parenti G, Carrozzo R, Coppa G, Felici L, Migliori V, et al. (July 1988). "X/Y translocation in a family with X-linked ichthyosis, chondrodysplasia punctata, and mental retardation: DNA analysis reveals deletion of the steroid sulphatase gene and translocation of its Y pseudogene".Clinical Genetics.34(1): 31–7.doi:10.1111/j.1399-0004.1988.tb02612.x.PMID3165728.S2CID39954996.
- Yen PH, Marsh B, Allen E, Tsai SP, Ellison J, Connolly L, et al. (December 1988). "The human X-linked steroid sulfatase gene and a Y-encoded pseudogene: evidence for an inversion of the Y chromosome during primate evolution".Cell.55(6): 1123–35.doi:10.1016/0092-8674(88)90257-7.PMID3203382.S2CID42065481.
- Chang PL, Varey PA, Rosa NE, Ameen M, Davidson RG (November 1986)."Association of steroid sulfatase with one of the arylsulfatase C isozymes in human fibroblasts".The Journal of Biological Chemistry.261(31): 14443–7.doi:10.1016/S0021-9258(18)66889-9.PMID3464600.
- Munroe DG, Chang PL (February 1987)."Tissue-specific expression of human arylsulfatase-C isozymes and steroid sulfatase".American Journal of Human Genetics.40(2): 102–14.PMC1684069.PMID3471087.
- Müller CR, Wahlström J, Ropers HH (1982). "Further evidence for the assignment of the steroid sulfatase X-linked ichthyosis locus to the telomer of Xp".Human Genetics.58(4): 446–447.doi:10.1007/bf00282842.PMID6948769.S2CID23372914.
- Migeon BR, Shapiro LJ, Norum RA, Mohandas T, Axelman J, Dabora RL (October 1982). "Differential expression of steroid sulphatase locus on active and inactive human X chromosome".Nature.299(5886): 838–40.Bibcode:1982Natur.299..838M.doi:10.1038/299838a0.PMID6957717.S2CID4361097.
- Bonaldo MF, Lennon G, Soares MB (September 1996)."Normalization and subtraction: two approaches to facilitate gene discovery".Genome Research.6(9): 791–806.doi:10.1101/gr.6.9.791.PMID8889548.
- Alperin ES, Shapiro LJ (August 1997)."Characterization of point mutations in patients with X-linked ichthyosis. Effects on the structure and function of the steroid sulfatase protein".The Journal of Biological Chemistry.272(33): 20756–63.doi:10.1074/jbc.272.33.20756.PMID9252398.
- Sugawara T, Shimizu H, Hoshi N, Fujimoto Y, Nakajima A, Fujimoto S (March 2000). "PCR diagnosis of X-linked ichthyosis: identification of a novel mutation (E560P) of the steroid sulfatase gene".Human Mutation.15(3): 296.doi:10.1002/(SICI)1098-1004(200003)15:3<296::AID-HUMU17>3.0.CO;2-#.PMID10679952.S2CID196599371.
- Oyama N, Satoh M, Iwatsuki K, Kaneko F (June 2000)."Novel point mutations in the steroid sulfatase gene in patients with X-linked ichthyosis: transfection analysis using the mutated genes".The Journal of Investigative Dermatology.114(6): 1195–9.doi:10.1046/j.1523-1747.2000.00004.x.PMID10844566.
- Jimenez Vaca AL, Valdes-Flores M, Rivera-Vega MR, González-Huerta LM, Kofman-Alfaro SH, Cuevas-Covarrubias SA (December 2001)."Deletion pattern of the STS gene in X-linked ichthyosis in a Mexican population".Molecular Medicine.7(12): 845–9.doi:10.1007/BF03401976.PMC1950010.PMID11844872.
- Hoffmann R, Rot A, Niiyama S, Billich A (December 2001)."Steroid sulfatase in the human hair follicle concentrates in the dermal papilla".The Journal of Investigative Dermatology.117(6): 1342–8.doi:10.1046/j.0022-202x.2001.01547.x.PMID11886493.
- Matsuoka R, Yanaihara A, Saito H, Furusawa Y, Toma Y, Shimizu Y, et al. (June 2002). "Regulation of estrogen activity in human endometrium: effect of IL-1β on steroid sulfatase activity in human endometrial stromal cells".Steroids.67(7): 655–9.doi:10.1016/S0039-128X(02)00016-8.PMID11996939.S2CID11872655.
External links[edit]
- Steryl-Sulfataseat the U.S. National Library of MedicineMedical Subject Headings(MeSH)