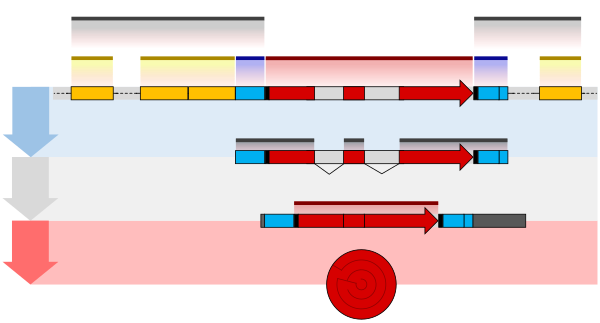
Regulatory sequence
Aregulatory sequenceis a segment of anucleic acidmolecule which is capable of increasing or decreasing theexpressionof specific genes within an organism.Regulation of gene expressionis an essential feature of all living organisms and viruses.
Description
[edit]|
|
InDNA,regulation ofgeneexpression normally happens at the level of RNA biosynthesis (transcription). It is accomplished through the sequence-specific binding of proteins (transcription factors) that activate or inhibit transcription. Transcription factors may act asactivators,repressors,or both. Repressors often act by preventingRNA polymerasefrom forming a productive complex with the transcriptional initiation region (promoter), while activators facilitate formation of a productive complex. Furthermore, DNA motifs have been shown to be predictive of epigenomic modifications, suggesting that transcription factors play a role in regulating theepigenome.[2]
InRNA,regulation may occur at the level of protein biosynthesis (translation), RNA cleavage,RNA splicing,or transcriptional termination. Regulatory sequences are frequently associated withmessenger RNA(mRNA) molecules, where they are used to control mRNA biogenesis or translation. A variety of biological molecules may bind to the RNA to accomplish this regulation, including proteins (e.g., translational repressors and splicing factors), other RNA molecules (e.g.,miRNA) andsmall molecules,in the case ofriboswitches.
Activation and implementation
[edit]A regulatory DNA sequence does not regulate unless it is activated. Different regulatory sequences are activated and then implement their regulation by different mechanisms.
Enhancer activation and implementation
[edit]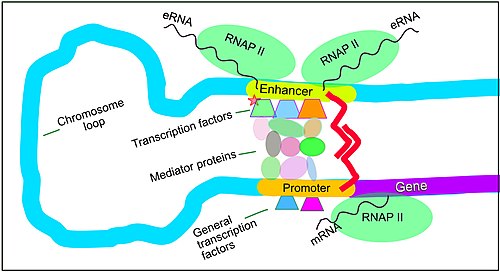
Expression of genes in mammals can be upregulated when signals are transmitted to the promoters associated with the genes.Cis-regulatory DNA sequencesthat are located in DNA regions distant from the promoters of genes can have very large effects on gene expression, with some genes undergoing up to 100-fold increased expression due to such acis-regulatory sequence.[3]Thesecis-regulatory sequences includeenhancers,silencers,insulatorsand tethering elements.[4]Among this constellation of sequences, enhancers and their associatedtranscription factor proteinshave a leading role in the regulation of gene expression.[5]
Enhancersare sequences of the genome that are major gene-regulatory elements. Enhancers control cell-type-specific gene expression programs, most often by looping through long distances to come in physical proximity with the promoters of their target genes.[6]In a study of brain cortical neurons, 24,937 loops were found, bringing enhancers to promoters.[3]Multiple enhancers, each often at tens or hundred of thousands of nucleotides distant from their target genes, loop to their target gene promoters and coordinate with each other to control expression of their common target gene.[6]
The schematic illustration in this section shows an enhancer looping around to come into close physical proximity with the promoter of a target gene. The loop is stabilized by a dimer of a connector protein (e.g. dimer ofCTCForYY1), with one member of the dimer anchored to its binding motif on the enhancer and the other member anchored to its binding motif on the promoter (represented by the red zigzags in the illustration).[7]Several cell function specific transcription factor proteins (in 2018 Lambert et al. indicated there were about 1,600 transcription factors in a human cell[8]) generally bind to specific motifs on an enhancer[9]and a small combination of these enhancer-bound transcription factors, when brought close to a promoter by a DNA loop, govern the level of transcription of the target gene.Mediator (coactivator)(a complex usually consisting of about 26 proteins in an interacting structure) communicates regulatory signals from enhancer DNA-bound transcription factors directly to the RNA polymerase II (RNAP II) enzyme bound to the promoter.[10]
Enhancers, when active, are generally transcribed from both strands of DNA with RNA polymerases acting in two different directions, producing two eRNAs as illustrated in the Figure.[11]An inactive enhancer may be bound by an inactive transcription factor. Phosphorylation of thetranscription factormay activate it and that activated transcription factor may then activate the enhancer to which it is bound (see small red star representing phosphorylation of a transcription factor bound to an enhancer in the illustration).[12]An activated enhancer begins transcription of its RNA before activating a promoter to initiate transcription of messengerRNAfrom its target gene.[13]
CpG island methylation and demethylation
[edit]
5-Methylcytosine(5-mC) is amethylatedform of theDNAbasecytosine(see figure). 5-mC is anepigeneticmarker found predominantly on cytosines within CpG dinucleotides, which consist of a cytosine is followed by a guanine reading in the 5′ to 3′ direction along the DNA strand (CpG sites). About 28 million CpG dinucleotides occur in the human genome.[14]In most tissues of mammals, on average, 70% to 80% of CpG cytosines are methylated (forming 5-methyl-CpG, or 5-mCpG).[15]Methylated cytosines within CpG sequences often occur in groups, calledCpG islands.About 59% of promoter sequences have a CpG island while only about 6% of enhancer sequences have a CpG island.[16]CpG islands constitute regulatory sequences, since if CpG islands are methylated in the promoter of a gene this can reduce or silence gene expression.[17]
DNA methylation regulates gene expression through interaction with methyl binding domain (MBD) proteins, such as MeCP2, MBD1 and MBD2. These MBD proteins bind most strongly to highly methylatedCpG islands.[18]These MBD proteins have both a methyl-CpG-binding domain and a transcriptional repression domain.[18]They bind to methylated DNA and guide or direct protein complexes with chromatin remodeling and/or histone modifying activity to methylated CpG islands. MBD proteins generally repress local chromatin by means such as catalyzing the introduction of repressive histone marks or creating an overall repressive chromatin environment throughnucleosomeremodeling and chromatin reorganization.[18]
Transcription factorsare proteins that bind to specific DNA sequences in order to regulate the expression of a given gene. The binding sequence for a transcription factor in DNA is usually about 10 or 11 nucleotides long. There are approximately 1,400 different transcription factors encoded in the human genome and they constitute about 6% of all human protein coding genes.[19]About 94% of transcription factor binding sites that are associated with signal-responsive genes occur in enhancers while only about 6% of such sites occur in promoters.[9]
EGR1is a transcription factor important for regulation of methylation of CpG islands. An EGR1 transcription factor binding site is frequently located in enhancer or promoter sequences.[20]There are about 12,000 binding sites for EGR1 in the mammalian genome and about half of EGR1 binding sites are located in promoters and half in enhancers.[20]The binding of EGR1 to its target DNA binding site is insensitive to cytosine methylation in the DNA.[20]
While only small amounts of EGR1 protein are detectable in cells that are un-stimulated, EGR1 translation into protein at one hour after stimulation is markedly elevated.[21]Expression of EGR1 in various types of cells can be stimulated by growth factors, neurotransmitters, hormones, stress and injury.[21]In the brain, when neurons are activated, EGR1 proteins are upregulated, and they bind to (recruit) pre-existing TET1 enzymes, which are highly expressed in neurons.TET enzymescan catalyze demethylation of 5-methylcytosine. When EGR1 transcription factors bring TET1 enzymes to EGR1 binding sites in promoters, the TET enzymes candemethylatethe methylated CpG islands at those promoters. Upon demethylation, these promoters can then initiate transcription of their target genes. Hundreds of genes in neurons are differentially expressed after neuron activation through EGR1 recruitment of TET1 to methylated regulatory sequences in their promoters.[20]
Activation by double- or single-strand breaks
[edit]About 600 regulatory sequences in promoters and about 800 regulatory sequences in enhancers appear to depend on double-strand breaks initiated bytopoisomerase 2β(TOP2B) for activation.[22][23]The induction of particular double-strand breaks is specific with respect to the inducing signal. When neurons are activatedin vitro,just 22 TOP2B-induced double-strand breaks occur in their genomes.[24]However, whencontextual fear conditioningis carried out in a mouse, this conditioning causes hundreds of gene-associated DSBs in the medial prefrontal cortex and hippocampus, which are important for learning and memory.[25]

Such TOP2B-induced double-strand breaks are accompanied by at least four enzymes of thenon-homologous end joining (NHEJ) DNA repair pathway(DNA-PKcs, KU70, KU80 and DNA LIGASE IV) (see figure). These enzymes repair the double-strand breaks within about 15 minutes to 2 hours.[24][26]The double-strand breaks in the promoter are thus associated with TOP2B and at least these four repair enzymes. These proteins are present simultaneously on a single promoter nucleosome (there are about 147 nucleotides in the DNA sequence wrapped around a single nucleosome) located near the transcription start site of their target gene.[26]
The double-strand break introduced by TOP2B apparently frees the part of the promoter at an RNA polymerase–bound transcription start site to physically move to its associated enhancer. This allows the enhancer, with its bound transcription factors and mediator proteins, to directly interact with the RNA polymerase that had been paused at the transcription start site to start transcription.[24][10]
Similarly, topoisomerase I (TOP1) enzymes appear to be located at many enhancers, and those enhancers become activated when TOP1 introduces a single-strand break.[27]TOP1 causes single-strand breaks in particular enhancer DNA regulatory sequences when signaled by a specific enhancer-binding transcription factor.[27]Topoisomerase I breaks are associated with different DNA repair factors than those surrounding TOP2B breaks. In the case of TOP1, the breaks are associated most immediately with DNA repair enzymesMRE11,RAD50andATR.[27]
Examples
[edit]Genomes can be analyzed systematically to identify regulatory regions.[28]Conserved non-coding sequencesoften contain regulatory regions, and so they are often the subject of these analyses.
- CAAT box
- CCAAT box
- Operator (biology)
- Pribnow box
- TATA box
- SECIS element,mRNA
- Polyadenylationsignal, mRNA
- A-box
- Z-box
- C-box
- E-box
- G-box
Insulin gene
[edit]Regulatory sequences for theinsulin geneare:[29]
- A5
- Z
- negative regulatory element(NRE)[30]
- C2
- E2
- A3
- cAMP response element
- A2
- CAAT enhancer binding(CEB)
- C1
- E1
- G1
See also
[edit]- Regulator gene
- Regulation of gene expression
- Cis-acting element
- Gene regulatory network
- Open Regulatory Annotation Database
- Operon
- DNA binding site
- Promoter
- Trans-acting factor
- ORegAnno
References
[edit]- ^abShafee, Thomas; Lowe, Rohan (2017)."Eukaryotic and prokaryotic gene structure".WikiJournal of Medicine.4(1).doi:10.15347/wjm/2017.002.ISSN2002-4436.
- ^Whitaker JW, Zhao Chen, Wei Wang. (2014) Predicting the Human Epigenome from DNA Motifs. Nature Methods. doi:10.1038/nmeth.3065
- ^abBeagan JA, Pastuzyn ED, Fernandez LR, Guo MH, Feng K, Titus KR, et al. (June 2020)."Three-dimensional genome restructuring across timescales of activity-induced neuronal gene expression".Nature Neuroscience.23(6): 707–717.doi:10.1038/s41593-020-0634-6.PMC7558717.PMID32451484.
- ^Verheul TC, van Hijfte L, Perenthaler E, Barakat TS (2020)."The Why of YY1: Mechanisms of Transcriptional Regulation by Yin Yang 1".Frontiers in Cell and Developmental Biology.8:592164.doi:10.3389/fcell.2020.592164.PMC7554316.PMID33102493.
- ^Spitz F, Furlong EE (September 2012). "Transcription factors: from enhancer binding to developmental control".Nature Reviews. Genetics.13(9): 613–26.doi:10.1038/nrg3207.PMID22868264.S2CID205485256.
- ^abSchoenfelder S, Fraser P (August 2019). "Long-range enhancer-promoter contacts in gene expression control".Nature Reviews. Genetics.20(8): 437–455.doi:10.1038/s41576-019-0128-0.PMID31086298.S2CID152283312.
- ^Weintraub AS, Li CH, Zamudio AV, Sigova AA, Hannett NM, Day DS, et al. (December 2017)."YY1 Is a Structural Regulator of Enhancer-Promoter Loops".Cell.171(7): 1573–1588.e28.doi:10.1016/j.cell.2017.11.008.PMC5785279.PMID29224777.
- ^Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, et al. (February 2018)."The Human Transcription Factors".Cell.172(4): 650–665.doi:10.1016/j.cell.2018.01.029.PMID29425488.
- ^abGrossman SR, Engreitz J, Ray JP, Nguyen TH, Hacohen N, Lander ES (July 2018)."Positional specificity of different transcription factor classes within enhancers".Proceedings of the National Academy of Sciences of the United States of America.115(30): E7222–E7230.doi:10.1073/pnas.1804663115.PMC6065035.PMID29987030.
- ^abAllen BL, Taatjes DJ (March 2015)."The Mediator complex: a central integrator of transcription".Nature Reviews. Molecular Cell Biology.16(3): 155–66.doi:10.1038/nrm3951.PMC4963239.PMID25693131.
- ^Mikhaylichenko O, Bondarenko V, Harnett D, Schor IE, Males M, Viales RR, Furlong EE (January 2018)."The degree of enhancer or promoter activity is reflected by the levels and directionality of eRNA transcription".Genes & Development.32(1): 42–57.doi:10.1101/gad.308619.117.PMC5828394.PMID29378788.
- ^Li QJ, Yang SH, Maeda Y, Sladek FM, Sharrocks AD, Martins-Green M (January 2003)."MAP kinase phosphorylation-dependent activation of Elk-1 leads to activation of the co-activator p300".The EMBO Journal.22(2): 281–91.doi:10.1093/emboj/cdg028.PMC140103.PMID12514134.
- ^Carullo NV, Phillips Iii RA, Simon RC, Soto SA, Hinds JE, Salisbury AJ, et al. (September 2020)."Enhancer RNAs predict enhancer-gene regulatory links and are critical for enhancer function in neuronal systems".Nucleic Acids Research.48(17): 9550–9570.doi:10.1093/nar/gkaa671.PMC7515708.PMID32810208.
- ^Lövkvist C, Dodd IB, Sneppen K, Haerter JO (June 2016)."DNA methylation in human epigenomes depends on local topology of CpG sites".Nucleic Acids Research.44(11): 5123–32.doi:10.1093/nar/gkw124.PMC4914085.PMID26932361.
- ^Jabbari K, Bernardi G (May 2004). "Cytosine methylation and CpG, TpG (CpA) and TpA frequencies".Gene.333:143–9.doi:10.1016/j.gene.2004.02.043.PMID15177689.
- ^Steinhaus R, Gonzalez T, Seelow D, Robinson PN (June 2020)."Pervasive and CpG-dependent promoter-like characteristics of transcribed enhancers".Nucleic Acids Research.48(10): 5306–5317.doi:10.1093/nar/gkaa223.PMC7261191.PMID32338759.
- ^Bird A (January 2002)."DNA methylation patterns and epigenetic memory".Genes & Development.16(1): 6–21.doi:10.1101/gad.947102.PMID11782440.
- ^abcDu Q, Luu PL, Stirzaker C, Clark SJ (2015)."Methyl-CpG-binding domain proteins: readers of the epigenome".Epigenomics.7(6): 1051–73.doi:10.2217/epi.15.39.PMID25927341.
- ^Vaquerizas JM, Kummerfeld SK, Teichmann SA, Luscombe NM (April 2009). "A census of human transcription factors: function, expression and evolution".Nature Reviews. Genetics.10(4): 252–63.doi:10.1038/nrg2538.PMID19274049.S2CID3207586.
- ^abcdSun Z, Xu X, He J, Murray A, Sun MA, Wei X, et al. (August 2019)."EGR1 recruits TET1 to shape the brain methylome during development and upon neuronal activity".Nature Communications.10(1): 3892.Bibcode:2019NatCo..10.3892S.doi:10.1038/s41467-019-11905-3.PMC6715719.PMID31467272.
- ^abKubosaki A, Tomaru Y, Tagami M, Arner E, Miura H, Suzuki T, et al. (2009)."Genome-wide investigation of in vivo EGR-1 binding sites in monocytic differentiation".Genome Biology.10(4): R41.doi:10.1186/gb-2009-10-4-r41.PMC2688932.PMID19374776.
- ^Dellino GI, Palluzzi F, Chiariello AM, Piccioni R, Bianco S, Furia L, et al. (June 2019)."Release of paused RNA polymerase II at specific loci favors DNA double-strand-break formation and promotes cancer translocations".Nature Genetics.51(6): 1011–1023.doi:10.1038/s41588-019-0421-z.PMID31110352.S2CID159041612.
- ^Singh S, Szlachta K, Manukyan A, Raimer HM, Dinda M, Bekiranov S, Wang YH (March 2020)."Pausing sites of RNA polymerase II on actively transcribed genes are enriched in DNA double-stranded breaks".J Biol Chem.295(12): 3990–4000.doi:10.1074/jbc.RA119.011665.PMC7086017.PMID32029477.
- ^abcMadabhushi R, Gao F, Pfenning AR, Pan L, Yamakawa S, Seo J, et al. (June 2015)."Activity-Induced DNA Breaks Govern the Expression of Neuronal Early-Response Genes".Cell.161(7): 1592–605.doi:10.1016/j.cell.2015.05.032.PMC4886855.PMID26052046.
- ^Stott RT, Kritsky O, Tsai LH (2021)."Profiling DNA break sites and transcriptional changes in response to contextual fear learning".PLOS ONE.16(7): e0249691.Bibcode:2021PLoSO..1649691S.doi:10.1371/journal.pone.0249691.PMC8248687.PMID34197463.
- ^abJu BG, Lunyak VV, Perissi V, Garcia-Bassets I, Rose DW, Glass CK, Rosenfeld MG (June 2006). "A topoisomerase IIbeta-mediated dsDNA break required for regulated transcription".Science.312(5781): 1798–802.Bibcode:2006Sci...312.1798J.doi:10.1126/science.1127196.PMID16794079.S2CID206508330.
- ^abcPuc J, Kozbial P, Li W, Tan Y, Liu Z, Suter T, et al. (January 2015)."Ligand-dependent enhancer activation regulated by topoisomerase-I activity".Cell.160(3): 367–80.doi:10.1016/j.cell.2014.12.023.PMC4422651.PMID25619691.
- ^Stepanova M, Tiazhelova T, Skoblov M, Baranova A (May 2005)."A comparative analysis of relative occurrence of transcription factor binding sites in vertebrate genomes and gene promoter areas".Bioinformatics.21(9): 1789–96.doi:10.1093/bioinformatics/bti307.PMID15699025.
- ^Melloul D, Marshak S, Cerasi E (March 2002)."Regulation of insulin gene transcription".Diabetologia.45(3): 309–26.doi:10.1007/s00125-001-0728-y.PMID11914736.
- ^Jang WG, Kim EJ, Park KG, Park YB, Choi HS, Kim HJ, et al. (January 2007). "Glucocorticoid receptor mediated repression of human insulin gene expression is regulated by PGC-1alpha".Biochemical and Biophysical Research Communications.352(3): 716–21.doi:10.1016/j.bbrc.2006.11.074.PMID17150186.