Chromosome 1
| Chromosome 1 | |
|---|---|
 Human chromosome 1 pair afterG-banding.One is from mother, one is from father. | |
 Chromosome 1 pair in human malekaryogram. | |
| Features | |
| Length (bp) | 248,387,328 bp (CHM13) |
| No.of genes | 1,961 (CCDS)[1] |
| Type | Autosome |
| Centromere position | Metacentric[2] (123.4 Mbp[3]) |
| Complete gene lists | |
| CCDS | Gene list |
| HGNC | Gene list |
| UniProt | Gene list |
| NCBI | Gene list |
| External map viewers | |
| Ensembl | Chromosome 1 |
| Entrez | Chromosome 1 |
| NCBI | Chromosome 1 |
| UCSC | Chromosome 1 |
| Full DNA sequences | |
| RefSeq | NC_000001(FASTA) |
| GenBank | CM000663(FASTA) |
Chromosome 1is the designation for the largesthuman chromosome.Humans have two copies of chromosome 1, as they do with all of theautosomes,which are the non-sex chromosomes.Chromosome 1 spans about 249 millionnucleotidebase pairs,which are the basic units of information forDNA.[4]It represents about 8% of the total DNA in human cells.[5]
It was the last completed chromosome, sequenced two decades after the beginning of theHuman Genome Project.
Genes
[edit]Number of genes
[edit]The following are some of the gene count estimates of human chromosome 1. Because researchers use different approaches togenome annotationtheir predictions of thenumber of geneson each chromosome varies (for technical details, seegene prediction). Among various projects, the collaborative consensus coding sequence project (CCDS) takes an extremely conservative strategy. So CCDS's gene number prediction represents a lower bound on the total number of human protein-coding genes.[6]
| Estimated by | Protein-coding genes | Non-coding RNA genes | Pseudogenes | Source | Release date |
|---|---|---|---|---|---|
| CCDS | 1,961 | — | — | [1] | 2016-09-08 |
| HGNC | 1,993 | 707 | 1,113 | [7] | 2017-05-12 |
| Ensembl | 2,044 | 1,924 | 1,223 | [8] | 2017-03-29 |
| UniProt | 2,064 | — | — | [9] | 2018-02-28 |
| NCBI | 2,093 | 1,790 | 1,426 | [10][11][12] | 2017-05-19 |
Gene list
[edit]The following is a partial list of genes on human chromosome 1. For complete list, see the link in the infobox on the right.
- C1orf112:encoding protein Chromosome 1 open reading frame 112
- C1orf127:encoding protein Chromosome 1 open reading frame 127
- C1orf27:encoding protein Chromosome 1 open reading frame 27
- C1orf38:encoding protein Chromosome 1 open reading frame 38
- CCDC181:encoding protein Coiled-coil domain-containing protein 181
- DENN1B:hypothesized to be related to asthma
- FHAD1:encoding protein Forkhead-associated domain containing protein 1
- LOC100132287:uncharacterized protein
- LRRIQ3:encoding protein Leucine-rich repeats and IQ motif containing 3
- Shisa family member 4:encoding protein Shisa family member 4
- TINAGL1:encoding protein Tubulointerstitial nephritis antigen-like
p-arm
[edit]Partial list of the genes located on p-arm (short arm) of human chromosome 1:
- AADACL3:Arylacetamide deacetylase-like 3
- AADACL4:Arylacetamide deacetylase-like 4
- ACADM:acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain
- ACTG1P6:encoding protein Actin, gamma 1 pseudogene 6
- ACTL8:Actin-like 8
- ADGRL2(1p31.1): adhesion G protein-coupled receptor L2
- ADPRHL2:Poly(ADP-ribose) glycohydrolase ARH3
- AMPD2:encodingenzymeAMP deaminase 2
- ARID1A(1p36)
- ATXN7L2:Ataxin 7-like 2
- AZIN2:encodingenzymeAntizyme inhibitor 2 (AzI2) also known as arginine decarboxylase (ADC)
- BCAS2:Breast carcinoma amplified sequence 2
- BCL10(1p22)
- LRIF1:encodingproteinLigand-dependent nuclear receptor-interacting factor 1
- C1orf109:chromosome 1 open reading frame 109
- C1orf162:encoding protein Chromosome 1 open reading frame 162
- C1orf194:encoding protein Chromosome 1 open reading frame 194
- CZIB:chromosome 1 open reading frame 123
- CACHD1encodingproteinCache domain containing 1
- CAMK2N1:encoding protein Calcium/calmodulin dependent protein kinase II inhibitor 1
- CAMTA1(1p36)
- CASP9(1p36)
- CASZ1(1p36): Castor zinc finger 1
- CCDC17:encoding protein Coiled-coil domain containing 17
- CCDC18:encoding protein Coiled-coil domain containing 18
- CEP85:encoding protein Centrosomal protein 85
- CFAP74:encoding protein Cilia and flagella associated protein 74
- CHD5(1p36)
- CLIC4(1p36)
- CLSPN(1p34)
- CMPK:UMP-CMP kinase
- COL16A1(1p35)
- COL11A1:collagen, type XI, Alpha 1
- CPT2:carnitine palmitoyltransferase II
- CRYZ:Crystallin zeta
- CSDE1:Cold shock domain containing E1
- CYP4B1(1p33)
- CYR61(1p22)
- DBT:dihydrolipoamide branched chain transacylase E2
- DCLRE1B:DNA cross-link repair 1B
- DEPDC1encodingproteinDEP domain containing 1
- DIRAS3(1p31): DIRAS family, GTP-binding RAS-like 3
- DISP3:encoding protein Dispatched rnd transporter family member 3
- DNASE2B:encoding protein Deoxyribonuclease 2 beta
- DPH5:Diphthine synthase
- DVL1(1p36)
- ENO1(1p36)
- EPHA2(1p36)
- EPS15(1p32)
- ESPN:espin (autosomal recessive deafness 36)
- EVI5:ecotropic viral integration site 5
- EXO5:encoding protein Exonuclease 5
- EXTL1:exostosin like glycosyltransferase 1
- EXTL2:exostosin like glycosyltransferase 2
- FAAH:Fatty-acid amide hydrolase 1
- FAM46B:family with sequence similarity 46, member B
- FAM46C:family with sequence similarity 46, member C
- FAM76A:family with sequence similarity 76, member A
- FAM87B:encoding protein Family with sequence similarity 87 member B
- FBXO2:F-box protein 2
- FNBP1LencodingproteinFormin-binding protein 1-like
- FPGT:Fucose-1-phosphate guanylyltransferase
- FUBP1(1p31)
- GALE:UDP-galactose-4-epimerase
- GADD45A(1p31)
- GBP1(1p22)
- GBP2:guanylate binding protein 2
- GBP5encodingproteinGuanylate binding protein 5
- GBP6:encoding protein Guanylate binding protein family member 6
- GJB3:gap junction protein, beta 3, 31kDa (connexin 31)
- GLMN(1p22)
- GNL2:G protein nucleolar 2
- GSTM1(1p13)
- GUCA2B:encoding protein Guanylate cyclase activator 2B
- HDAC1(1p35)
- HES2:Hes family bHLH transcription factor 2
- HES3:Hes family bHLH transcription factor 3
- HMGCL:3-hydroxymethyl-3-methylglutaryl-Coenzyme A lyase (hydroxymethylglutaricaciduria)
- HAO2encodingproteinHydroxyacid oxidase 2
- HMGCS2:3-hydroxy-3-methylglutaryl-CoA synthase 2
- HP1BP3:Heterochromatin protein 1, binding protein 3
- IFI6:Interferon Alpha -inducible protein 6
- IL22RA1(1p36)
- INTS11:Integrator complex subunit 11
- JAK1(1p31)
- JUN(1p32)
- KANK4:encoding protein KN motif and ankyrin repeat domains 4
- KCNQ4:potassium voltage-gated channel, KQT-like subfamily, member 4
- KIF1B:kinesin family member 1B
- L1TD1:LINE-1 type transposase domain containing 1
- LCK(1p35)
- LINC01137:encoding protein Long intergenic non-protein coding RNA 1137
- LRRC39:Leucine-rich repeat-containing protein 39
- LRRC40:Leucine-rich repeat-containing protein 40
- LRRC41:Leucine-rich repeat-containing protein 41
- LRRC8D:Leucine-rich repeat-containing protein 8D
- MACO1:encoding protein Transmembrane protein 57
- MAN1A2:Mannosyl-oligosaccharide 1,2- Alpha -mannosidase IB
- MAP3K6:encoding protein Mitogen-activated protein kinase kinase kinase 6
- MEAF6:MYST/ESA1 associated factor 6
- MECR:Trans-2-enoyl-CoA reductase, mitochondrial
- MFAP2:Microfibrillar-associated protein 2
- MIB2(1p36)
- MIER1(1p31)
- MIGA1:encoding protein Mitoguardin 1
- MFN2:mitofusin 2
- MFSD2:Major facilitator superfamily domain containing 2A
- MIR6079:microRNA 6079
- MMEL1:Membrane metallo-endopeptidase-like 1
- MTFR1L:mitochondrial fission regulator 1 like
- MTHFR(1p36): 5,10-methylenetetrahydrofolate reductase (NADPH)
- MUL1:Mitochondrial E3 ubiquitin protein ligase 1
- MUTYH(1p34): mutY homolog (E. coli)
- NBPF3:Neuroblastoma breakpoint family member 3
- NDUFA4P1:encoding protein Nadh dehydrogenase (ubiquinone) 1 Alpha subcomplex, 4, 9kda, pseudogene 1
- NGF:Nerve Growth Factor
- NOL9:Nucleolar protein 9
- NRAS(1p13)
- NOTCH2(1p12)
- OLFML3:Olfactomedin-like 3
- OMA1:Metalloendopeptidase OMA1, mitochondrial
- OVGP1:Oviductal glycoprotein 1
- PARK7(1p36): Parkinson disease (autosomal recessive, early onset) 7
- PINK1:PTEN induced putative kinase 1
- PLOD1:procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1
- PRAMEF10:encoding protein Prame family member 10
- PRMT6:Protein arginine methyltransferase 6
- PRXL2B:encoding protein Peroxiredoxin like 2B
- PSRC1:Proline/serine-rich coiled-coil protein 1
- RAD54L:RAD54-like
- RAP1A(1p13)
- RBM15(1p13)
- RCC2:Regulator of chromosome condensation 2
- REG4(1p12)
- RHBDL2:Rhomboid like 2
- RHOC(1p13)
- RIMS3:encoding protein Regulating synaptic membrane exocytosis 3
- RLF:rearranged L-myc fusion
- RNF11(1p32)
- RNF19B:encoding protein Ring finger protein 19B
- RNF220:RING finger protein 220
- RPA2(1p35)
- RSPO1(1p34)
- S100A1(1q21)
- SAMD11:encoding protein Sterile Alpha motif domain containing 11
- SDC3:Syndecan-3
- SDHB(1p36)
- SFPQ(1p34): encoding protein Splicing factor proline and glutamine rich
- SGIP1:SH3 domain GRB2-like protein 3-interaction protein 1
- SH3BGRL3:SH3 domain-binding glutamic acid-rich-like protein 3
- SLC16A1(1p13)
- SLC2A1Glucose transporter 1
- SPSB1:SPRY domain-containing SOCS box protein 1
- STIL(1p33)
- SYCP1:Synaptonemal complex protein 1
- SZT2:Seizure threshold 2 homolog
- TACSTD2:tumor-associated calcium signal transducer 2
- TAL1(1p33)
- TCTEX1D4:encoding protein Tctex1 domain containing 4
- TCEB3:Transcription elongation factor B polypeptide 3
- TGFBR3(1p22)
- THRAP3(1p34)
- TIE1(1p34)
- TM2D1:encoding protein Tm2 domain containing 1
- TMCO2:encodingproteintransmembrane and coiled-coil domains 2
- TMCO4:encodingproteintransmembrane and coiled-coil domains 4
- TMEM48:encodingproteinnucleoporinNDC1
- TMEM50A:Transmembrane protein 50A
- TMEM59:Transmembrane protein 59
- TMEM69:Transmembrane protein 69
- TMEM201encodingproteinTransmembrane protein 201
- TMEM222:Transmembrane protein 222
- TOE1:Target of EGR1 protein 1
- TRABD2B:encoding protein Trab domain containing 2b
- TRAPPC3:Trafficking protein particle complex subunit 3
- TRIT1:tRNA isopentenyltransferase, mitochondrial
- TSHB:thyroid stimulating hormone, beta
- TTC39A:Tetratricopeptide repeat 39A
- UBR4:E3 ubiquitin-protein ligase component n-recognin 4
- UROD:uroporphyrinogen decarboxylase (the gene forporphyria cutanea tarda)
- USP1(1p31)
- USP48:Ubiquitin carboxyl-terminal hydrolase 48
- VAV3(1p13)
- VPS13D:Vacuolar protein sorting-associated protein 13D
- VTCN1(1p13)
- WARS2:Tryptophanyl-tRNA synthetase, mitochondrial
- WDR77(1p13)
- YBX1(1p34)
- ZCCHC17:zinc finger CCHC-type containing 17
- ZFP69:encoding protein Zfp69 zinc finger protein
- ZMYM1encodingproteinZinc finger MYM-type containing 1
- ZNF436:Zinc finger protein 436
- ZNF684:encoding protein Zinc finger protein 684
- ZYG11BencodingproteinZyg-11 family member B, cell cycle regulator
- ZZZ3:ZZ-type zinc finger-containing protein 3
q-arm
[edit]Partial list of the genes located onq-arm(long arm) of human chromosome 1:
- ABL2(1q25)
- ADIPOR1(1q32)
- AHCTF1:encodingproteinELYS
- AKT3(1q43-44)
- ANGPTL1:Angiopoietin-related protein 1
- ARHGEF2(1q22)
- ARID4B:encodingproteinAT-rich interactive domain-containing protein 4B
- ARV1encodingproteinARV1 homolog (S. cerevisiae)
- ARNT(1q21)
- ASPM(1q31): a brain size determinant
- ATF3(1q32)
- ATP2B4(1q32)
- BCL9(1q21)
- CATSPERE:encoding protein Catsper channel auxiliary subunit epsilon
- C1orf21:chromosome 1 open reading frame 21
- MMTAP2encodingproteinMultiple myeloma tumor-associated protein 2
- TEX35:TEX35
- C1orf74:chromosome 1 open reading frame 74
- C2CD4D:C2 calcium-dependent domain-containing 4D
- CTSS:Cathepsin S
- CD5L:CD5 molecule like
- CENPL:Centromere protein L
- CENPF(1q41)
- CHTOP:Chromatin target of prmt1
- CNIH4:cornichon homolog 4
- CNST:Consortin
- CREG1:Cellular repressor of E1A stimulated genes 1
- CRP:C-reactive protein
- CRTC2(1q21)
- CSRP1:Cysteine and glycine rich protein 1
- DCAF8:encoding protein DDB1 and CUL4 associated factor 8
- DDX59:DEAD-box helicase 59
- DEL1Q21:encoding protein Chromosome 1q21.1 deletion syndrome
- DPT:Dermatopontin
- DISC2,long non-coding RNA
- DNAH14encodingproteinDynein, axonemal, heavy chain 14
- DUSP10(1q41)
- DUSP27:encoding protein Dual specificity phosphatase 27 (putative)
- ECM1(1q21)
- EDEM3:ER degradation enhancing Alpha -mannosidase like protein 3
- EGLN1(1q42)
- ELF3:encoding protein E74 like ets transcription factor 3
- ENAH(1q42)
- ESRRG(1q41)
- FAM129A:family with sequence similarity 129, member A
- FAM163A:encoding protein neuroblastoma-derived secretory protein
- FAM20B:FAM20B, glycosaminoglycan xylosylkinase
- FAM63A:Family with sequence similarity 63, member A
- FAM78B:family with sequence similarity 78, member B
- FAM89A:encoding protein Fam89A
- FBXO28:F-box protein 28
- FCMR:Fc fragment of IgM receptor
- FCGR2B(1q23)
- FCGR2C:encoding protein Fc fragment of igg receptor iic (gene/pseudogene)
- FH(1q43):fumarase
- FLAD1:encoding protein Flavin adenine dinucleotide synthetase 1
- FLG-AS1:encoding protein FLG antisense RNA 1
- FMO3:flavin containing monooxygenase 3
- FRA1JencodingproteinFragile site, 5-azacytidine type, common, fra(1)(q12)
- G0S2:encoding G0/G1 switch 2
- GAS5(1q25)
- GBA:glucosidase, beta; acid (includes glucosylceramidase) (gene forGaucher disease)
- GBAP1:glucosylceramidase beta pseudogene 1
- GLC1A:gene forglaucoma
- GON4L:gon-4 like
- GPA33(1q24)
- GPR37L1G protein-coupled receptor 37 like 1
- H3C13:encoding protein Histone cluster 2 h3 family member d
- HEATR1:HEAT repeat-containing protein 1
- HFE2:hemochromatosis type 2 (juvenile)
- HIST2H2AB:Histone 2A type 2-B
- HIST2H2BF:Histone H2B type 2-F
- HIST2H3PS2:Histone cluster 2, H3, pseudogene 2
- HIST3H2A:Histone H2A type 3
- HIST3H2BB:Histone H2B type 3-B
- HRM2:Hair, curly
- IGSF8(1q23)
- INAVA:Innate immunity activator protein
- INTS3:Integrator complex subunit 3
- IRF2BP2:encoding protein Interferon regulatory factor 2 binding protein 2
- IRF6:gene forconnective tissueformation
- KCNH1(1q32)
- KIF14(1q32)
- LEFTY1:Left-right determination factor 1
- LHX9encodingproteinLIM homeobox 9
- LMNA:lamin A/C
- LMOD1:encoding protein Leiomodin 1
- LOC645166encodingproteinLymphocyte-specific protein 1 pseudogene
- LYPLAL1:Lysophospholipase-like 1
- MAPKAPK2(1q32)
- MIR194-1:microRNA 194-1
- MIR5008:microRNA 5008
- MPC2:Mitochondrial pyruvate carrier 2
- MOSC1:MOCO sulphurase C-terminal domain containing 1
- MOSC2:MOSC domain-containing protein 2, mitochondrial
- MPZ:myelin protein zero (Charcot–Marie–Tooth neuropathy 1B)
- MSTO1:misato 1
- MTR:5-methyltetrahydrofolate-homocysteine methyltransferase
- NAV1:Neuron navigator 1
- NBPF16:Neuroblastoma breakpoint family, member 16
- NOC2L:Nucleolar complex protein 2 homolog
- NUCKS1:Nuclear ubiquitous casein and cyclin-dependent kinases substrate
- NVL:Nuclear valosin-containing protein-like
- OLFML2B:Olfactomedin-like 2B
- OPTC:Opticin
- OTUD7B:OTU domain-containing protein 7B
- PACERRencodingproteinPTGS2 antisense NFKB1 complex-mediated expression regulator RNA
- PBX1(1q23)
- PEA15(1q23)
- PGDB5:PiggyBac transposable element derived 5
- PIAS3(1q21)
- PI4KB:Phosphatidylinositol 4-kinase beta
- PIP5K1A(1q21): Phosphatidylinositol-4-phosphate 5-kinase type-1 Alpha
- PLA2G4A(1q31)
- PPOX:protoporphyrinogen oxidase
- PRCC(1q23)
- PRR9encodingproteinProline rich 9
- PSEN2(1q42): presenilin 2 (Alzheimer disease 4)
- PTGS2(1q31)
- PTPN14(1q32-41)
- PTPN7(1q32)
- RABIF:RAB interacting factor
- RASSF5(1q32)
- RGS2(1q31)
- RN5S1@:RNA, 5S ribosomal 1q42 cluster
- RPS27(1q21)
- SCAMP3:Secretory carrier-associated membrane protein 3
- SDHC(1q23)
- SELE(1q24)
- SFT2D2:encoding protein Sft2 domain containing 2
- SHC1(1q21)
- SHCBP1L:encoding protein Shc binding and spindle associated 1 like
- SLC30A10:encoding protein Solute carrier family 30 member 10
- SLC39A1(1q21)
- SLC50A1:Solute carrier family 50 member 1
- SMCP:Sperm mitochondrial-associated cysteine-rich protein
- SMG7:nonsense mediated mRNA decay factor
- SMYD3(1q44)
- SPG23
- SPRR1A:Cornifin-A
- SPRR1B:Cornifin-B
- SPRR2A:Small proline rich protein 2A
- SPRTN:Spartan
- TARBP1:TAR (HIV-1) RNA-binding protein 1
- TBCE:Tubulin-specific chaperone E
- THBS3:Thrombospondin 3
- TMCO1:Transmembrane and coiled-coil domain-containing protein 1
- TMEM9:Transmembrane protein 9
- TMEM63A:Transmembrane protein 63A
- TMEM81:Transmembrane protein 81
- TNFAIP8L2:encoding TNF Alpha induced protein 8 like 2
- TNFSF18(1q25)
- TNNT2:cardiac troponin T2
- TOR1AIP1:Torsin-1A-interacting protein 1
- TOR3A:encoding protein Torsin family 3 member A
- TP53BP2(1q41)
- TRE-CTC1-5:Transfer RNA-Glu (CTC) 1-5
- UAP1:UDP-N-acetylhexosamine pyrophosphorylase
- USH2A:Usher syndrome2A (autosomal recessive, mild)
- USF1(1q23)
- VANGL2:encoding protein VANGL planar cell polarity protein 2
- VPS45:Vacuolar protein sorting-associated protein 45
- VPS72:Vacuolar protein sorting-associated protein 72
- YY1AP1:YY1-associated protein 1
- ZBED6:zinc finger, BED-type containing 6
- ZC3H11A:Zinc finger CCCH domain-containing protein 11A
- ZNF648encodingproteinZinc finger protein 648
- ZNF669:Zinc finger protein 669
- ZNF687:Zinc finger protein 687
- ZNF692:Zinc finger protein 692
- ZNF695:Zinc finger protein 695
Diseases and disorders
[edit]There are 890 known diseases related to this chromosome.[citation needed]Some of these diseases arehearing loss,Alzheimer's disease,glaucomaandbreast cancer.Rearrangements and mutations of chromosome 1 are prevalent in cancer and many other diseases. Patterns of sequence variation reveal signals of recent selection in specific genes that may contribute to human fitness, and also in regions where no function is evident.
Completemonosomy(only having one copy of the entire chromosome) is invariably lethal before birth.[13]Completetrisomy(having three copies of the entire chromosome) is lethal within days afterconception.[13]Some partial deletions and partial duplications producebirth defects.
The following diseases are some of those related to genes on chromosome 1 (which contains the most knowngenetic diseasesof any human chromosome):
- 1q21.1 deletion syndrome
- 1q21.1 duplication syndrome
- Alzheimer's disease
- Autosomal dominant Charcot-Marie-Tooth disease type 2 with giant axons
- Bipolar disorder
- Breast cancer
- Brooke GreenbergDisease (Syndrome X)
- Carnitine palmitoyltransferase II deficiency
- Charcot–Marie–Tooth disease,types 1 and 2
- collagenopathy, types II and XI
- congenital hypothyroidism
- Ehlers-Danlos syndrome
- Factor V Leiden thrombophilia
- Familial adenomatous polyposis
- galactosemia
- Gaucher disease
- Gaucher-like disease
- Gelatinous drop-like corneal dystrophy
- Glaucoma
- GLUT1 deficiency syndrome
- Hearing loss,autosomal recessive deafness 36
- Hemochromatosis
- Hepatoerythropoietic porphyria
- Homocystinuria
- Hutchinson Gilford progeria syndrome
- 3-hydroxy-3-methylglutaryl-CoA lyase deficiency
- Hypertrophic cardiomyopathy,autosomal dominant mutations of TNNT2; hypertrophy usually mild; restrictive phenotype may be present; may carry high risk of sudden cardiac death
- maple syrup urine disease
- medium-chain acyl-coenzyme A dehydrogenase deficiency
- Microcephaly
- Muckle–Wells syndrome
- Nonsyndromic deafness
- Oligodendroglioma
- Parkinson disease
- Pheochromocytoma
- porphyria
- porphyria cutanea tarda
- popliteal pterygium syndrome
- prostate cancer
- Stickler syndrome
- TAR syndrome
- trimethylaminuria
- Usher syndrome
- Usher syndrome type II
- Van der Woude syndrome
- Variegate porphyria
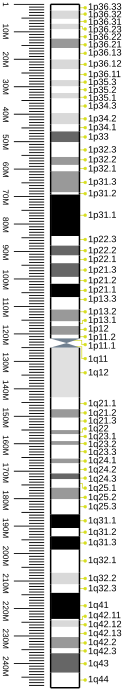
Cytogenetic band
[edit]| Chr. | Arm[18] | Band[19] | ISCN start[20] |
ISCN stop[20] |
Basepair start |
Basepair stop |
Stain[21] | Density |
|---|---|---|---|---|---|---|---|---|
| 1 | p | 36.33 | 0 | 100 | 1 | 2,300,000 | gneg | |
| 1 | p | 36.32 | 100 | 244 | 2,300,001 | 5,300,000 | gpos | 25 |
| 1 | p | 36.31 | 244 | 344 | 5,300,001 | 7,100,000 | gneg | |
| 1 | p | 36.23 | 344 | 459 | 7,100,001 | 9,100,000 | gpos | 25 |
| 1 | p | 36.22 | 459 | 660 | 9,100,001 | 12,500,000 | gneg | |
| 1 | p | 36.21 | 660 | 861 | 12,500,001 | 15,900,000 | gpos | 50 |
| 1 | p | 36.13 | 861 | 1206 | 15,900,001 | 20,100,000 | gneg | |
| 1 | p | 36.12 | 1206 | 1321 | 20,100,001 | 23,600,000 | gpos | 25 |
| 1 | p | 36.11 | 1321 | 1521 | 23,600,001 | 27,600,000 | gneg | |
| 1 | p | 35.3 | 1521 | 1651 | 27,600,001 | 29,900,000 | gpos | 25 |
| 1 | p | 35.2 | 1651 | 1780 | 29,900,001 | 32,300,000 | gneg | |
| 1 | p | 35.1 | 1780 | 1895 | 32,300,001 | 34,300,000 | gpos | 25 |
| 1 | p | 34.3 | 1895 | 2210 | 34,300,001 | 39,600,000 | gneg | |
| 1 | p | 34.2 | 2210 | 2411 | 39,600,001 | 43,700,000 | gpos | 25 |
| 1 | p | 34.1 | 2411 | 2770 | 43,700,001 | 46,300,000 | gneg | |
| 1 | p | 33 | 2770 | 2986 | 46,300,001 | 50,200,000 | gpos | 75 |
| 1 | p | 32.3 | 2986 | 3273 | 50,200,001 | 55,600,000 | gneg | |
| 1 | p | 32.2 | 3273 | 3416 | 55,600,001 | 58,500,000 | gpos | 50 |
| 1 | p | 32.1 | 3416 | 3732 | 58,500,001 | 60,800,000 | gneg | |
| 1 | p | 31.3 | 3732 | 3976 | 60,800,001 | 68,500,000 | gpos | 50 |
| 1 | p | 31.2 | 3976 | 4206 | 68,500,001 | 69,300,000 | gneg | |
| 1 | p | 31.1 | 4206 | 4852 | 69,300,001 | 84,400,000 | gpos | 100 |
| 1 | p | 22.3 | 4852 | 5210 | 84,400,001 | 87,900,000 | gneg | |
| 1 | p | 22.2 | 5210 | 5440 | 87,900,001 | 91,500,000 | gpos | 75 |
| 1 | p | 22.1 | 5440 | 5741 | 91,500,001 | 94,300,000 | gneg | |
| 1 | p | 21.3 | 5741 | 5957 | 94,300,001 | 99,300,000 | gpos | 75 |
| 1 | p | 21.2 | 5957 | 6029 | 99,300,001 | 101,800,000 | gneg | |
| 1 | p | 21.1 | 6029 | 6244 | 101,800,001 | 106,700,000 | gpos | 100 |
| 1 | p | 13.3 | 6244 | 6459 | 106,700,001 | 111,200,000 | gneg | |
| 1 | p | 13.2 | 6459 | 6660 | 111,200,001 | 115,500,000 | gpos | 50 |
| 1 | p | 13.1 | 6660 | 6861 | 115,500,001 | 117,200,000 | gneg | |
| 1 | p | 12 | 6861 | 7048 | 117,200,001 | 120,400,000 | gpos | 50 |
| 1 | p | 11.2 | 7048 | 7119 | 120,400,001 | 121,700,000 | gneg | |
| 1 | p | 11.1 | 7119 | 7335 | 121,700,001 | 123,400,000 | acen | |
| 1 | q | 11 | 7335 | 7579 | 123,400,001 | 125,100,000 | acen | |
| 1 | q | 12 | 7579 | 8483 | 125,100,001 | 143,200,000 | gvar | |
| 1 | q | 21.1 | 8483 | 8756 | 143,200,001 | 147,500,000 | gneg | |
| 1 | q | 21.2 | 8756 | 8957 | 147,500,001 | 150,600,000 | gpos | 50 |
| 1 | q | 21.3 | 8957 | 9244 | 150,600,001 | 155,100,000 | gneg | |
| 1 | q | 22 | 9244 | 9459 | 155,100,001 | 156,600,000 | gpos | 50 |
| 1 | q | 23.1 | 9459 | 9832 | 156,600,001 | 159,100,000 | gneg | |
| 1 | q | 23.2 | 9832 | 10048 | 159,100,001 | 160,500,000 | gpos | 50 |
| 1 | q | 23.3 | 10048 | 10349 | 160,500,001 | 165,500,000 | gneg | |
| 1 | q | 24.1 | 10349 | 10507 | 165,500,001 | 167,200,000 | gpos | 50 |
| 1 | q | 24.2 | 10507 | 10679 | 167,200,001 | 170,900,000 | gneg | |
| 1 | q | 24.3 | 10679 | 10894 | 170,900,001 | 173,000,000 | gpos | 75 |
| 1 | q | 25.1 | 10894 | 11009 | 173,000,001 | 176,100,000 | gneg | |
| 1 | q | 25.2 | 11009 | 11196 | 176,100,001 | 180,300,000 | gpos | 50 |
| 1 | q | 25.3 | 11196 | 11598 | 180,300,001 | 185,800,000 | gneg | |
| 1 | q | 31.1 | 11598 | 11827 | 185,800,001 | 190,800,000 | gpos | 100 |
| 1 | q | 31.2 | 11827 | 11942 | 190,800,001 | 193,800,000 | gneg | |
| 1 | q | 31.3 | 11942 | 12172 | 193,800,001 | 198,700,000 | gpos | 100 |
| 1 | q | 32.1 | 12172 | 12617 | 198,700,001 | 207,100,000 | gneg | |
| 1 | q | 32.2 | 12617 | 12803 | 207,100,001 | 211,300,000 | gpos | 25 |
| 1 | q | 32.3 | 12803 | 13033 | 211,300,001 | 214,400,000 | gneg | |
| 1 | q | 41 | 13033 | 13320 | 214,400,001 | 223,900,000 | gpos | 100 |
| 1 | q | 42.11 | 13320 | 13406 | 223,900,001 | 224,400,000 | gneg | |
| 1 | q | 42.12 | 13406 | 13607 | 224,400,001 | 226,800,000 | gpos | 25 |
| 1 | q | 42.13 | 13607 | 13966 | 226,800,001 | 230,500,000 | gneg | |
| 1 | q | 42.2 | 13966 | 14153 | 230,500,001 | 234,600,000 | gpos | 50 |
| 1 | q | 42.3 | 14153 | 14397 | 234,600,001 | 236,400,000 | gneg | |
| 1 | q | 43 | 14397 | 14756 | 236,400,001 | 243,500,000 | gpos | 75 |
| 1 | q | 44 | 14756 | 15100 | 243,500,001 | 248,956,422 | gneg |
References
[edit]- ^ab"Search results - 1[CHR] AND" Homo sapiens "[Organism] AND (" has ccds "[Properties] AND alive[prop]) - Gene".NCBI.CCDS Release 20 forHomo sapiens.2016-09-08.Retrieved2017-05-28.
- ^Tom Strachan; Andrew Read (2 April 2010).Human Molecular Genetics.Garland Science. p. 45.ISBN978-1-136-84407-2.
- ^abcGenome Decoration Page, NCBI.Ideogram data for Homo sapience (850 bphs, Assembly GRCh38.p3).Last update 2014-06-03. Retrieved 2017-04-26.
- ^http://vega.sanger.ac.uk/Homo_sapiens/mapview?chr=1Chromosome size and number of genes derived from this database, retrieved 2012-03-11.
- ^Gregory SG, Barlow KF, McLay KE, Kaul R, Swarbreck D, Dunham A, et al. (May 2006)."The DNA sequence and biological annotation of human chromosome 1".Nature.441(7091): 315–21.Bibcode:2006Natur.441..315G.doi:10.1038/nature04727.PMID16710414.
- ^Pertea M, Salzberg SL (2010)."Between a chicken and a grape: estimating the number of human genes".Genome Biol.11(5): 206.doi:10.1186/gb-2010-11-5-206.PMC2898077.PMID20441615.
- ^"Statistics & Downloads for chromosome 1".HUGO Gene Nomenclature Committee.2017-05-12. Archived fromthe originalon 2017-06-29.Retrieved2017-05-19.
- ^"Chromosome 1: Chromosome summary - Homo sapiens".Ensembl Release 88.2017-03-29.Retrieved2017-05-19.
- ^"Human chromosome 1: entries, gene names and cross-references to MIM".UniProt.2018-02-28.Retrieved2018-03-16.
- ^"Search results - 1[CHR] AND" Homo sapiens "[Organism] AND (" genetype protein coding "[Properties] AND alive[prop]) - Gene".NCBI.2017-05-19.Retrieved2017-05-20.
- ^"Search results - 1[CHR] AND" Homo sapiens "[Organism] AND ( (" genetype miscrna "[Properties] OR" genetype ncrna "[Properties] OR" genetype rrna "[Properties] OR" genetype trna "[Properties] OR" genetype scrna "[Properties] OR" genetype snrna "[Properties] OR" genetype snorna "[Properties]) NOT" genetype protein coding "[Properties] AND alive[prop]) - Gene".NCBI.2017-05-19.Retrieved2017-05-20.
- ^"Search results - 1[CHR] AND" Homo sapiens "[Organism] AND (" genetype pseudo "[Properties] AND alive[prop]) - Gene".NCBI.2017-05-19.Retrieved2017-05-20.
- ^abGersen, Steven L.; Keagle, Martha B. (2013-03-26).The Principles of Clinical Cytogenetics.Springer Science & Business Media. p. 278.ISBN9781441916884.
- ^Genome Decoration Page, NCBI.Ideogram data for Homo sapience (400 bphs, Assembly GRCh38.p3).Last update 2014-03-04. Retrieved 2017-04-26.
- ^Genome Decoration Page, NCBI.Ideogram data for Homo sapience (550 bphs, Assembly GRCh38.p3).Last update 2015-08-11. Retrieved 2017-04-26.
- ^International Standing Committee on Human Cytogenetic Nomenclature (2013).ISCN 2013: An International System for Human Cytogenetic Nomenclature (2013).Karger Medical and Scientific Publishers.ISBN978-3-318-02253-7.
- ^Sethakulvichai, W.; Manitpornsut, S.; Wiboonrat, M.; Lilakiatsakun, W.; Assawamakin, A.; Tongsima, S. (2012)."Estimation of band level resolutions of human chromosome images".2012 Ninth International Conference on Computer Science and Software Engineering (JCSSE).pp. 276–282.doi:10.1109/JCSSE.2012.6261965.ISBN978-1-4673-1921-8.S2CID16666470.
- ^"p":Short arm;"q":Long arm.
- ^For cytogenetic banding nomenclature, see articlelocus.
- ^abThese values (ISCN start/stop) are based on the length of bands/ideograms from the ISCN book, An International System for Human Cytogenetic Nomenclature (2013).Arbitrary unit.
- ^gpos:Region which is positively stained byG banding,generallyAT-richand gene poor;gneg:Region which is negatively stained by G banding, generallyCG-richand gene rich;acenCentromere.var:Variable region;stalk:Stalk.
Further reading
[edit]- Murphy WJ, Fronicke L, O'Brien SJ, Stanyon R (2003)."The Origin of Human Chromosome 1 and Its Homologs in Placental Mammals".Genome Res.13(8): 1880–8.doi:10.1101/gr.1022303.PMC403779.PMID12869576.
- Revera, M.; Van Der Merwe, L; et al. (2007)."Long-term follow-up of R403WMHY7 and R92WTNNT2 HCM families: mutations determine left ventricular dimensions but not wall thickness during disease progression".Cardiovascular Journal of Africa.18(3): 146–153.PMC4213759.PMID17612745.
- Schutte BC, Carpten JD, Forus A, Gregory SG, Horii A, White PS (2001). "Report and abstracts of the sixth international workshop on human chromosome 1 mapping 2000. Iowa City, Iowa, USA. 30 September-3 October 2000".Cytogenet Cell Genet.92(1–2): 23–41.doi:10.1159/000056867.PMID11306795.S2CID202651814.
External links
[edit]- National Institutes of Health."Chromosome 1".Genetics Home Reference.Archived fromthe originalon 2012-02-04.Retrieved2017-05-06.
- "Final genome 'chapter' published".BBC News.2006-05-18.Retrieved2017-05-06.
- "Chromosome 1".Human Genome Project Information Archive 1990–2003.Retrieved2017-05-06.