Gene
| Part of a series on |
| Genetics |
|---|
 |
Inbiology,the wordgenehas two meanings. The Mendelian gene is a basic unit ofheredity.The molecular gene is a sequence ofnucleotidesinDNAthat is transcribed to produce a functionalRNA.There are two types of molecular genes: protein-coding genes and non-coding genes.[1][2][3]
Duringgene expression(the synthesis ofRNA or proteinfrom a gene), DNA is firstcopied into RNA.RNA can bedirectly functionalor be the intermediatetemplatefor the synthesis of a protein.
The transmission of genes to an organism'soffspring,is the basis of the inheritance ofphenotypic traitsfrom one generation to the next. These genes make up different DNA sequences, together called agenotype,that is specific to every given individual, within thegene poolof apopulationof a givenspecies.The genotype, along with environmental and developmental factors, ultimately determines thephenotypeof the individual.
Most biological traits occur under the combined influence ofpolygenes(a set of different genes) andgene–environment interactions.Some genetic traits are instantly visible, such aseye coloror the number of limbs, others are not, such asblood type,the risk for specific diseases, or the thousands of basicbiochemicalprocesses that constitutelife.
A gene can acquiremutationsin itssequence,leading to different variants, known asalleles,in thepopulation.These alleles encode slightly different versions of a gene, which may cause differentphenotypicaltraits.[4]Genesevolvedue tonatural selectionorsurvival of the fittestandgenetic driftof the alleles.
The termgenewas introduced by Danish botanist, plant physiologist, and geneticistWilhelm Johannsenin 1909.[5]It is inspired by theancient Greek:γόνος,gonos,that means offspring and procreation.
Definitions
[edit]There are many different ways to use the term "gene" based on different aspects of their inheritance, selection, biological function, or molecular structure but most of these definitions fall into two categories, the Mendelian gene or the molecular gene.[1][6][7][8][9]
The Mendelian gene is the classical gene of genetics and it refers to any heritable trait. This is the gene described inThe Selfish Gene.[10]More thorough discussions of this version of a gene can be found in the articlesGeneticsandGene-centered view of evolution.
The molecular gene definition is more commonly used across biochemistry, molecular biology, and most of genetics — the gene that is described in terms of DNA sequence.[1]There are many different definitions of this gene — some of which are misleading or incorrect.[6][11]
Very early work in the field that becamemolecular geneticssuggested the concept thatone gene makes one protein(originally 'one gene - one enzyme').[12][13]However, genes that produce repressor RNAs were proposed in the 1950s[14]and by the 1960s, textbooks were using molecular gene definitions that included those that specified functional RNA molecules such as ribosomal RNA and tRNA (noncoding genes) as well as protein-coding genes.[15]
This idea of two kinds of genes is still part of the definition of a gene in most textbooks. For example,
The primary function of the genome is to produce RNA molecules. Selected portions of the DNA nucleotide sequence are copied into a corresponding RNA nucleotide sequence, which either encodes a protein (if it is an mRNA) or forms a 'structural' RNA, such as a transfer RNA (tRNA) or ribosomal RNA (rRNA) molecule. Each region of the DNA helix that produces a functional RNA molecule constitutes a gene.[16]
We define a gene as a DNA sequence that is transcribed. This definition includes genes that do not encode proteins (not all transcripts are messenger RNA). The definition normally excludes regions of the genome that control transcription but are not themselves transcribed. We will encounter some exceptions to our definition of a gene - surprisingly, there is no definition that is entirely satisfactory.[17]
A gene is a DNA sequence that codes for a diffusible product. This product may be protein (as is the case in the majority of genes) or may be RNA (as is the case of genes that code for tRNA and rRNA). The crucial feature is that the product diffuses away from its site of synthesis to act elsewhere.[18]
The important parts of such definitions are: (1) that a gene corresponds to a transcription unit; (2) that genes produce both mRNA and noncoding RNAs; and (3) regulatory sequences control gene expression but are not part of the gene itself. However, there's one other important part of the definition and it is emphasized in Kostas Kampourakis' bookMaking Sense of Genes.
Therefore in this book I will consider genes as DNA sequences encoding information for functional products, be it proteins or RNA molecules. With 'encoding information', I mean that the DNA sequence is used as a template for the production of an RNA molecule or a protein that performs some function.[6]
The emphasis on function is essential because there are stretches of DNA that produce non-functional transcripts and they do not qualify as genes. These include obvious examples such as transcribed pseudogenes as well as less obvious examples such as junk RNA produced as noise due to transcription errors. In order to qualify as a true gene, by this definition, one has to prove that the transcript has a biological function.[6]
Early speculations on the size of a typical gene were based on high-resolution genetic mapping and on the size of proteins and RNA molecules. A length of 1500 base pairs seemed reasonable at the time (1965).[15]This was based on the idea that the gene was the DNA that was directly responsible for production of the functional product. The discovery of introns in the 1970s meant that many eukaryotic genes were much larger than the size of the functional product would imply. Typical mammalian protein-coding genes, for example, are about 62,000 base pairs in length (transcribed region) and since there are about 20,000 of them they occupy about 35–40% of the mammalian genome (including the human genome).[19][20][21]
In spite of the fact that both protein-coding genes and noncoding genes have been known for more than 50 years, there are still a number of textbooks, websites, and scientific publications that define a gene as a DNA sequence that specifies a protein. In other words, the definition is restricted to protein-coding genes. Here is an example from a recent article in American Scientist.
... to truly assess the potential significance of de novo genes, we relied on a strict definition of the word "gene" with which nearly every expert can agree. First, in order for a nucleotide sequence to be considered a true gene, an open reading frame (ORF) must be present. The ORF can be thought of as the "gene itself"; it begins with a starting mark common for every gene and ends with one of three possible finish line signals. One of the key enzymes in this process, the RNA polymerase, zips along the strand of DNA like a train on a monorail, transcribing it into its messenger RNA form. This point brings us to our second important criterion: A true gene is one that is both transcribed and translated. That is, a true gene is first used as a template to make transient messenger RNA, which is then translated into a protein.[22]
This restricted definition is so common that it has spawned many recent articles that criticize this "standard definition" and call for a new expanded definition that includes noncoding genes.[23][24][25]However, this so-called "new" definition has been around for more than half a century and it is not clear why some modern writers are ignoring noncoding genes.[editorializing]
Although some definitions can be more broadly applicable than others, the fundamental complexity of biology means that no definition of a gene can capture all aspects perfectly. Not all genomes are DNA (e.g.RNA viruses),[26]bacterialoperonsare multiple protein-coding regions transcribed into single large mRNAs,alternative splicingenables a single genomic region to encode multiple district products andtrans-splicingconcatenates mRNAs from shorter coding sequence across the genome.[27][28][29]Since molecular definitions exclude elements such as introns, promotors, and otherregulatory regions,these are instead thought of as "associated" with the gene and affect its function.
An even broader operational definition is sometimes used to encompass the complexity of these diverse phenomena, where a gene is defined as a union of genomic sequences encoding a coherent set of potentially overlapping functional products.[30]This definition categorizes genes by their functional products (proteins or RNA) rather than their specific DNA loci, with regulatory elements classified asgene-associatedregions.[30]
History
[edit]Discovery of discrete inherited units
[edit]
The existence of discrete inheritable units was first suggested byGregor Mendel(1822–1884).[31]From 1857 to 1864, inBrno,Austrian Empire(today's Czech Republic), he studied inheritance patterns in 8000 common ediblepea plants,tracking distinct traits from parent to offspring. He described these mathematically as 2ncombinations where n is the number of differing characteristics in the original peas. Although he did not use the termgene,he explained his results in terms of discrete inherited units that give rise to observable physical characteristics. This description prefiguredWilhelm Johannsen's distinction betweengenotype(the genetic material of an organism) andphenotype(the observable traits of that organism). Mendel was also the first to demonstrateindependent assortment,the distinction betweendominantandrecessivetraits, the distinction between aheterozygoteandhomozygote,and the phenomenon of discontinuous inheritance.
Prior to Mendel's work, the dominant theory of heredity was one ofblending inheritance,[32]which suggested that each parent contributed fluids to the fertilization process and that the traits of the parents blended and mixed to produce the offspring.Charles Darwindeveloped a theory of inheritance he termedpangenesis,fromGreekpan ( "all, whole" ) and genesis ( "birth" ) / genos ( "origin" ).[33][34]Darwin used the termgemmuleto describe hypothetical particles that would mix during reproduction.
Mendel's work went largely unnoticed after its first publication in 1866, but was rediscovered in the late 19th century byHugo de Vries,Carl Correns,andErich von Tschermak,who (claimed to have) reached similar conclusions in their own research.[35]Specifically, in 1889, Hugo de Vries published his bookIntracellular Pangenesis,[36]in which he postulated that different characters have individual hereditary carriers and that inheritance of specific traits in organisms comes in particles. De Vries called these units "pangenes" (Pangensin German), after Darwin's 1868 pangenesis theory.
Twenty years later, in 1909,Wilhelm Johannsenintroduced the term "gene"[5]and, in 1906,William Bateson,that of "genetics"[37][30]whileEduard Strasburger,among others, still used the term "pangene" for the fundamental physical and functional unit of heredity.[36]: Translator's preface, viii
Discovery of DNA
[edit]Advances in understanding genes and inheritance continued throughout the 20th century.Deoxyribonucleic acid(DNA) was shown to be the molecular repository of genetic information by experiments in the 1940s to 1950s.[38][39]The structure of DNA was studied byRosalind FranklinandMaurice WilkinsusingX-ray crystallography,which ledJames D. WatsonandFrancis Crickto publish a model of the double-stranded DNA molecule whose pairednucleotide basesindicated a compelling hypothesis for the mechanism of genetic replication.[40][41]
In the early 1950s the prevailing view was that the genes in a chromosome acted like discrete entities arranged like beads on a string. The experiments ofBenzerusingmutantsdefective in therII region of bacteriophage T4(1955–1959) showed that individual genes have a simple linear structure and are likely to be equivalent to a linear section of DNA.[42][43]
Collectively, this body of research established thecentral dogma of molecular biology,which states thatproteinsare translated fromRNA,which is transcribed fromDNA.This dogma has since been shown to have exceptions, such asreverse transcriptioninretroviruses.The modern study ofgeneticsat the level of DNA is known asmolecular genetics.
In 1972,Walter Fiersand his team were the first to determine the sequence of a gene: that ofbacteriophage MS2coat protein.[44]The subsequent development ofchain-terminationDNA sequencingin 1977 byFrederick Sangerimproved the efficiency of sequencing and turned it into a routine laboratory tool.[45]An automated version of the Sanger method was used in early phases of theHuman Genome Project.[46]
Modern synthesis and its successors
[edit]The theories developed in the early 20th century to integrateMendelian geneticswithDarwinian evolutionare called themodern synthesis,a term introduced byJulian Huxley.[47]
This view of evolution was emphasized byGeorge C. Williams'gene-centric view of evolution.He proposed that the Mendelian gene is aunitofnatural selectionwith the definition: "that which segregates and recombines with appreciable frequency."[48]: 24 Related ideas emphasizing the centrality of Mendelian genes and the importance of natural selection in evolution were popularized byRichard Dawkins.[10][49]
The development of theneutral theory of evolutionin the late 1960s led to the recognition that random genetic drift is a major player in evolution and that neutral theory should be the null hypothesis of molecular evolution.[50]This led to the construction ofphylogenetic treesand the development of themolecular clock,which is the basis of all dating techniques using DNA sequences. These techniques are not confined to molecular gene sequences but can be used on all DNA segments in the genome.
Molecular basis
[edit]
DNA
[edit]The vast majority of organisms encode their genes in long strands ofDNA(deoxyribonucleic acid). DNA consists of achainmade from four types ofnucleotidesubunits, each composed of: a five-carbon sugar (2-deoxyribose), aphosphategroup, and one of the fourbasesadenine,cytosine,guanine,andthymine.[51]: 2.1
Two chains of DNA twist around each other to form a DNAdouble helixwith the phosphate–sugar backbone spiraling around the outside, and the bases pointing inward with adeninebase pairingto thymine and guanine to cytosine. The specificity of base pairing occurs because adenine and thymine align to form twohydrogen bonds,whereas cytosine and guanine form three hydrogen bonds. The two strands in a double helix must, therefore, becomplementary,with their sequence of bases matching such that the adenines of one strand are paired with the thymines of the other strand, and so on.[51]: 4.1
Due to the chemical composition of thepentoseresidues of the bases, DNA strands have directionality. One end of a DNApolymercontains an exposedhydroxylgroup on thedeoxyribose;this is known as the3' endof the molecule. The other end contains an exposedphosphategroup; this is the5' end.The two strands of a double-helix run in opposite directions. Nucleic acid synthesis, includingDNA replicationandtranscriptionoccurs in the 5'→3' direction, because new nucleotides are added via adehydration reactionthat uses the exposed 3' hydroxyl as anucleophile.[52]: 27.2
Theexpressionof genes encoded in DNA begins bytranscribingthe gene intoRNA,a second type of nucleic acid that is very similar to DNA, but whose monomers contain the sugarriboserather thandeoxyribose.RNA also contains the baseuracilin place ofthymine.RNA molecules are less stable than DNA and are typically single-stranded. Genes that encode proteins are composed of a series of three-nucleotidesequences calledcodons,which serve as the "words" in the genetic "language". Thegenetic codespecifies the correspondence duringprotein translationbetween codons andamino acids.The genetic code is nearly the same for all known organisms.[51]: 4.1
Chromosomes
[edit]

The total complement of genes in an organism or cell is known as itsgenome,which may be stored on one or morechromosomes.A chromosome consists of a single, very long DNA helix on which thousands of genes are encoded.[51]: 4.2 The region of the chromosome at which a particular gene is located is called itslocus.Each locus contains onealleleof a gene; however, members of a population may have different alleles at the locus, each with a slightly different gene sequence.
The majority ofeukaryoticgenes are stored on a set of large, linear chromosomes. The chromosomes are packed within thenucleusin complex with storage proteins calledhistonesto form a unit called anucleosome.DNA packaged and condensed in this way is calledchromatin.[51]: 4.2 The manner in which DNA is stored on the histones, as well as chemical modifications of the histone itself, regulate whether a particular region of DNA is accessible forgene expression.In addition to genes, eukaryotic chromosomes contain sequences involved in ensuring that the DNA is copied without degradation of end regions and sorted into daughter cells during cell division:replication origins,telomeres,and thecentromere.[51]: 4.2 Replication origins are the sequence regions whereDNA replicationis initiated to make two copies of the chromosome. Telomeres are long stretches of repetitive sequences that cap the ends of the linear chromosomes and prevent degradation of coding and regulatory regions duringDNA replication.The length of the telomeres decreases each time the genome is replicated and has been implicated in theagingprocess.[54]The centromere is required for bindingspindle fibresto separate sister chromatids into daughter cells duringcell division.[51]: 18.2
Prokaryotes(bacteriaandarchaea) typically store their genomes on a single, large,circular chromosome.Similarly, some eukaryoticorganellescontain a remnant circular chromosome with a small number of genes.[51]: 14.4 Prokaryotes sometimes supplement their chromosome with additional small circles of DNA calledplasmids,which usually encode only a few genes and are transferable between individuals. For example, the genes forantibiotic resistanceare usually encoded on bacterial plasmids and can be passed between individual cells, even those of different species, viahorizontal gene transfer.[55]
Whereas the chromosomes of prokaryotes are relatively gene-dense, those of eukaryotes often contain regions of DNA that serve no obvious function. Simple single-celled eukaryotes have relatively small amounts of such DNA, whereas the genomes of complexmulticellular organisms,including humans, contain an absolute majority of DNA without an identified function.[56]This DNA has often been referred to as "junk DNA".However, more recent analyses suggest that, although protein-coding DNA makes up barely 2% of thehuman genome,about 80% of the bases in the genome may be expressed, so the term "junk DNA" may be a misnomer.[27]
Structure and function
[edit]Structure
[edit]|
|
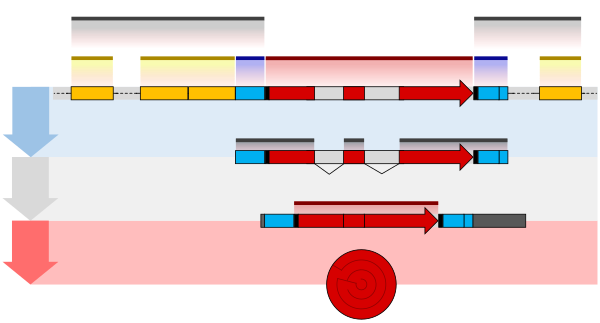
Thestructure of a protein-coding geneconsists of many elements of which the actualprotein coding sequenceis often only a small part. These include introns and untranslated regions of the mature mRNA. Noncoding genes can also contain introns that are removed during processing to produce the mature functional RNA.
All genes are associated withregulatory sequencesthat are required for their expression. First, genes require apromotersequence. The promoter is recognized and bound bytranscription factorsthat recruit and helpRNA polymerasebind to the region to initiate transcription.[51]: 7.1 The recognition typically occurs as aconsensus sequencelike theTATA box.A gene can have more than one promoter, resulting in messenger RNAs (mRNA) that differ in how far they extend in the 5' end.[58]Highly transcribed genes have "strong" promoter sequences that form strong associations with transcription factors, thereby initiating transcription at a high rate. Others genes have "weak" promoters that form weak associations with transcription factors and initiate transcription less frequently.[51]: 7.2 Eukaryoticpromoterregions are much more complex and difficult to identify thanprokaryoticpromoters.[51]: 7.3
Additionally, genes can have regulatory regions many kilobases upstream or downstream of the gene that alter expression. These act bybindingto transcription factors which then cause the DNA to loop so that the regulatory sequence (and bound transcription factor) become close to the RNA polymerase binding site.[59]For example,enhancersincrease transcription by binding anactivatorprotein which then helps to recruit the RNA polymerase to the promoter; converselysilencersbindrepressorproteins and make the DNA less available for RNA polymerase.[60]
The mature messenger RNA produced from protein-coding genes containsuntranslated regionsat both ends which contain binding sites forribosomes,RNA-binding proteins,miRNA,as well asterminator,andstartandstop codons.[61]In addition, most eukaryoticopen reading framescontain untranslatedintrons,which are removed andexons,which are connected together in a process known asRNA splicing.Finally, the ends of gene transcripts are defined bycleavage and polyadenylation (CPA) sites,where newly produced pre-mRNA gets cleaved and a string of ~200 adenosine monophosphates is added at the 3' end. Thepoly(A)tail protects mature mRNA from degradation and has other functions, affecting translation, localization, and transport of the transcript from the nucleus. Splicing, followed by CPA, generate the finalmature mRNA,which encodes the protein or RNA product.[62]
Many noncoding genes in eukaryotes have different transcription termination mechanisms and they do not have poly(A) tails.
Many prokaryotic genes are organized intooperons,with multiple protein-coding sequences that are transcribed as a unit.[63][64]The genes in anoperonare transcribed as a continuousmessenger RNA,referred to as apolycistronic mRNA.The termcistronin this context is equivalent to gene. The transcription of an operon's mRNA is often controlled by arepressorthat can occur in an active or inactive state depending on the presence of specific metabolites.[65]When active, the repressor binds to a DNA sequence at the beginning of the operon, called theoperator region,and repressestranscriptionof theoperon;when the repressor is inactive transcription of the operon can occur (see e.g.Lac operon). The products of operon genes typically have related functions and are involved in the sameregulatory network.[51]: 7.3
Complexity
[edit]Though many genes have simple structures, as with much of biology, others can be quite complex or represent unusual edge-cases. Eukaryotic genes often have introns are often much larger than their exons,[66][67]and those introns can even have other genesnested inside them.[68]Associated enhancers may be many kilobase away, or even on entirely different chromosomes operating via physical contact between two chromosomes.[69][70]A single gene can encode multiple different functional products byalternative splicing,and conversely gene may be split across chromosomes but those transcripts are concatenated back together into a functional sequence bytrans-splicing.[71]It is also possible foroverlapping genesto share some of their DNA sequence, either on opposite strands or the same strand (in a different reading frame, or even the same reading frame).[72]
Gene expression
[edit]In all organisms, two steps are required to read the information encoded in a gene's DNA and produce the protein it specifies. First, the gene's DNA istranscribedto messenger RNA (mRNA).[51]: 6.1 Second, that mRNA istranslatedto protein.[51]: 6.2 RNA-coding genes must still go through the first step, but are not translated into protein.[73]The process of producing a biologically functional molecule of either RNA or protein is calledgene expression,and the resulting molecule is called agene product.
Genetic code
[edit]
The nucleotide sequence of a gene's DNA specifies the amino acid sequence of a protein through thegenetic code.Sets of three nucleotides, known ascodons,each correspond to a specific amino acid.[51]: 6 The principle that three sequential bases of DNA code for each amino acid was demonstrated in 1961 using frameshift mutations in the rIIB gene of bacteriophage T4[74](seeCrick, Brenner et al. experiment).
Additionally, a "start codon",and three"stop codons"indicate the beginning and end of theprotein coding region.There are 64 possible codons (four possible nucleotides at each of three positions, hence 43possible codons) and only 20 standard amino acids; hence the code is redundant and multiple codons can specify the same amino acid. The correspondence between codons and amino acids is nearly universal among all known living organisms.[75]
Transcription
[edit]Transcriptionproduces a single-strandedRNAmolecule known asmessenger RNA,whose nucleotide sequence is complementary to the DNA from which it was transcribed.[51]: 6.1 The mRNA acts as an intermediate between the DNA gene and its final protein product. The gene's DNA is used as a template to generate acomplementarymRNA. The mRNA matches the sequence of the gene's DNAcoding strandbecause it is synthesised as the complement of thetemplate strand.Transcription is performed by anenzymecalled anRNA polymerase,which reads the template strand in the3'to5'direction and synthesizes the RNA from5'to3'.To initiate transcription, the polymerase first recognizes and binds apromoterregion of the gene. Thus, a major mechanism ofgene regulationis the blocking or sequestering the promoter region, either by tight binding byrepressormolecules that physically block the polymerase or by organizing the DNA so that the promoter region is not accessible.[51]: 7
Inprokaryotes,transcription occurs in thecytoplasm;for very long transcripts, translation may begin at the 5' end of the RNA while the 3' end is still being transcribed. Ineukaryotes,transcription occurs in the nucleus, where the cell's DNA is stored. The RNA molecule produced by the polymerase is known as theprimary transcriptand undergoespost-transcriptional modificationsbefore being exported to the cytoplasm for translation. One of the modifications performed is thesplicingofintronswhich are sequences in the transcribed region that do not encode a protein.Alternative splicingmechanisms can result in mature transcripts from the same gene having different sequences and thus coding for different proteins. This is a major form of regulation in eukaryotic cells and also occurs in some prokaryotes.[51]: 7.5 [76]
Translation
[edit]
Translationis the process by which amature mRNAmolecule is used as a template for synthesizing a newprotein.[51]: 6.2 Translation is carried out byribosomes,large complexes of RNA and protein responsible for carrying out the chemical reactions to add newamino acidsto a growingpolypeptide chainby the formation ofpeptide bonds.The genetic code is read three nucleotides at a time, in units calledcodons,via interactions with specialized RNA molecules calledtransfer RNA(tRNA). Each tRNA has three unpaired bases known as theanticodonthat are complementary to the codon it reads on the mRNA. The tRNA is alsocovalentlyattached to theamino acidspecified by the complementary codon. When the tRNA binds to its complementary codon in an mRNA strand, the ribosome attaches its amino acid cargo to the new polypeptide chain, which is synthesized fromamino terminustocarboxyl terminus.During and after synthesis, most new proteins mustfoldto their activethree-dimensional structurebefore they can carry out their cellular functions.[51]: 3
Regulation
[edit]Genes are regulatedso that they areexpressedonly when the product is needed, since expression draws on limited resources.[51]: 7 A cell regulates its gene expression depending on itsexternal environment(e.g.available nutrients,temperatureand otherstresses), its internal environment (e.g.cell division cycle,metabolism,infection status), and itsspecific roleif in amulticellularorganism. Gene expression can be regulated at any step: fromtranscriptional initiation,toRNA processing,topost-translational modificationof the protein. The regulation oflactosemetabolism genes inE. coli(lacoperon) was the first such mechanism to be described in 1961.[77]
RNA genes
[edit]A typical protein-coding gene is first copied intoRNAas an intermediate in the manufacture of the final protein product.[51]: 6.1 In other cases, the RNA molecules are the actual functional products, as in the synthesis ofribosomal RNAandtransfer RNA.Some RNAs known asribozymesare capable ofenzymatic function,while others such asmicroRNAsandriboswitcheshave regulatory roles. TheDNAsequences from which such RNAs are transcribed are known asnon-coding RNA genes.[73]
Somevirusesstore their entire genomes in the form ofRNA,and contain no DNA at all.[78][79]Because they use RNA to store genes, theircellularhostsmay synthesize their proteins as soon as they areinfectedand without the delay in waiting for transcription.[80]On the other hand, RNAretroviruses,such asHIV,require thereverse transcriptionof theirgenomefrom RNA into DNA before their proteins can be synthesized.
Inheritance
[edit]
Organisms inherit their genes from their parents.Asexualorganisms simply inherit a complete copy of their parent's genome.Sexualorganisms have two copies of each chromosome because they inherit one complete set from each parent.[51]: 1
Mendelian inheritance
[edit]According toMendelian inheritance,variations in an organism'sphenotype(observable physical and behavioral characteristics) are due in part to variations in itsgenotype(particular set of genes). Each gene specifies a particular trait with a different sequence of a gene (alleles) giving rise to different phenotypes. Most eukaryotic organisms (such as the pea plants Mendel worked on) have two alleles for each trait, one inherited from each parent.[51]: 20
Alleles at a locus may bedominantorrecessive;dominant alleles give rise to their corresponding phenotypes when paired with any other allele for the same trait, whereas recessive alleles give rise to their corresponding phenotype only when paired with another copy of the same allele. If you know the genotypes of the organisms, you can determine which alleles are dominant and which are recessive. For example, if the allele specifying tall stems in pea plants is dominant over the allele specifying short stems, then pea plants that inherit one tall allele from one parent and one short allele from the other parent will also have tall stems. Mendel's work demonstrated that alleles assort independently in the production ofgametes,orgerm cells,ensuring variation in the next generation. Although Mendelian inheritance remains a good model for many traits determined by single genes (including a number of well-knowngenetic disorders) it does not include the physical processes of DNA replication and cell division.[81][82]
DNA replication and cell division
[edit]The growth, development, and reproduction of organisms relies oncell division;the process by which a singlecelldivides into two usually identicaldaughter cells.This requires first making a duplicate copy of every gene in thegenomein a process calledDNA replication.[51]: 5.2 The copies are made by specializedenzymesknown asDNA polymerases,which "reads" one strand of the double-helical DNA, known as the template strand, and synthesize a new complementary strand. Because the DNA double helix is held together bybase pairing,the sequence of one strand completely specifies the sequence of its complement; hence only one strand needs to be read by the enzyme to produce a faithful copy. The process of DNA replication issemiconservative;that is, the copy of the genome inherited by each daughter cell contains one original and one newly synthesized strand of DNA.[51]: 5.2
The rate of DNA replication in living cells was first measured as the rate of phage T4 DNA elongation in phage-infectedE. coliand found to be impressively rapid.[83]During the period of exponential DNA increase at 37 °C, the rate of elongation was 749 nucleotides per second.
After DNA replication is complete, the cell must physically separate the two copies of the genome and divide into two distinct membrane-bound cells.[51]: 18.2 Inprokaryotes(bacteriaandarchaea) this usually occurs via a relatively simple process calledbinary fission,in which each circular genome attaches to thecell membraneand is separated into the daughter cells as the membraneinvaginatesto split thecytoplasminto two membrane-bound portions. Binary fission is extremely fast compared to the rates of cell division ineukaryotes.Eukaryotic cell division is a more complex process known as thecell cycle;DNA replication occurs during a phase of this cycle known asS phase,whereas the process of segregatingchromosomesand splitting thecytoplasmoccurs duringM phase.[51]: 18.1
Molecular inheritance
[edit]The duplication and transmission of genetic material from one generation of cells to the next is the basis for molecular inheritance and the link between the classical and molecular pictures of genes. Organisms inherit the characteristics of their parents because the cells of the offspring contain copies of the genes in their parents' cells. Inasexually reproducingorganisms, the offspring will be a genetic copy orcloneof the parent organism. Insexually reproducingorganisms, a specialized form of cell division calledmeiosisproduces cells calledgametesorgerm cellsthat arehaploid,or contain only one copy of each gene.[51]: 20.2 The gametes produced by females are calledeggsor ova, and those produced by males are calledsperm.Two gametes fuse to form adiploidfertilized egg,a single cell that has two sets of genes, with one copy of each gene from the mother and one from the father.[51]: 20
During the process of meiotic cell division, an event calledgenetic recombinationorcrossing-overcan sometimes occur, in which a length of DNA on onechromatidis swapped with a length of DNA on the corresponding homologous non-sister chromatid. This can result in reassortment of otherwise linked alleles.[51]: 5.5 The Mendelian principle of independent assortment asserts that each of a parent's two genes for each trait will sort independently into gametes; which allele an organism inherits for one trait is unrelated to which allele it inherits for another trait. This is in fact only true for genes that do not reside on the same chromosome or are located very far from one another on the same chromosome. The closer two genes lie on the same chromosome, the more closely they will be associated in gametes and the more often they will appear together (known asgenetic linkage).[84]Genes that are very close are essentially never separated because it is extremely unlikely that a crossover point will occur between them.[84]
Molecular evolution
[edit]Mutation
[edit]DNA replication is for the most part extremely accurate, however errors (mutations) do occur.[51]: 7.6 The error rate ineukaryoticcellscan be as low as 10−8pernucleotideper replication,[85][86]whereas for some RNA viruses it can be as high as 10−3.[87]This means that each generation, each human genome accumulates around 30 new mutations.[88]Small mutations can be caused byDNA replicationand the aftermath ofDNA damageand includepoint mutationsin which a single base is altered andframeshift mutationsin which a single base is inserted or deleted. Either of these mutations can change the gene bymissense(change acodonto encode a different amino acid) ornonsense(a prematurestop codon).[89]Larger mutations can be caused by errors in recombination to causechromosomal abnormalitiesincluding theduplication,deletion, rearrangement or inversion of large sections of a chromosome. Additionally, DNA repair mechanisms can introduce mutational errors when repairing physical damage to the molecule. The repair, even with mutation, is more important to survival than restoring an exact copy, for example when repairingdouble-strand breaks.[51]: 5.4
When multiple differentallelesfor a gene are present in a species's population it is calledpolymorphic.Most different alleles are functionally equivalent, however some alleles can give rise to differentphenotypic traits.A gene's most common allele is called thewild type,and rare alleles are calledmutants.Thegenetic variationin relative frequencies of different alleles in a population is due to bothnatural selectionandgenetic drift.[90]The wild-type allele is not necessarily theancestorof less common alleles, nor is it necessarilyfitter.
Most mutations within genes areneutral,having no effect on the organism's phenotype (silent mutations). Some mutations do not change the amino acid sequence because multiple codons encode the same amino acid (synonymous mutations). Other mutations can be neutral if they lead to amino acid sequence changes, but the protein still functions similarly with the new amino acid (e.g.conservative mutations). Many mutations, however, aredeleteriousor evenlethal,and are removed from populations by natural selection. Genetic disorders are the result of deleterious mutations and can be due to spontaneous mutation in the affected individual, or can be inherited. Finally, a small fraction of mutations arebeneficial,improving the organism'sfitnessand are extremely important for evolution, since theirdirectional selectionleads to adaptiveevolution.[51]: 7.6
Sequence homology
[edit]The relationship between genes can be measured by comparing thesequencesof their DNA. If the level of similarity exceeds a minimum value, one can conclude that the genes descend from a common ancestor; they arehomologous.[91][92]Genes that are related by direct descent from a common ancestor are orthologous genes - they are usually found at the same locus in different species. Genes that are related as a result of a gene duplication event are parologous genes.[93][94]
It is often assumed that the functions of orthologous genes are more similar than those of paralogous genes, although the difference is minimal.[95][96]
Origins of new genes
[edit]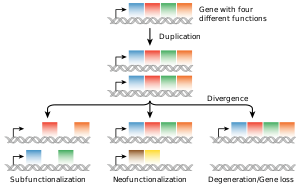
The most common source of new genes in eukaryotic lineages isgene duplication,which createscopy number variationof an existing gene in the genome.[97][98]The resulting genes (paralogs) may then diverge in sequence and in function. Sets of genes formed in this way compose agene family.Gene duplications and losses within a family are common and represent a major source of evolutionarybiodiversity.[99]Sometimes, gene duplication may result in a nonfunctional copy of a gene, or a functional copy may be subject to mutations that result in loss of function; such nonfunctional genes are calledpseudogenes.[51]: 7.6
"Orphan" genes,whose sequence shows no similarity to existing genes, are less common than gene duplicates. The human genome contains an estimate 18[100]to 60[101]genes with no identifiable homologs outside humans. Orphan genes arise primarily from eitherde novoemergencefrom previouslynon-coding sequence,or gene duplication followed by such rapid sequence change that the original relationship becomes undetectable.[102]De novogenes are typically shorter and simpler in structure than most eukaryotic genes, with few if any introns.[97]Over long evolutionary time periods,de novogene birth may be responsible for a significant fraction of taxonomically restricted gene families.[103]
Horizontal gene transferrefers to the transfer of genetic material through a mechanism other thanreproduction.This mechanism is a common source of new genes inprokaryotes,sometimes thought to contribute more to genetic variation than gene duplication.[104]It is a common means of spreadingantibiotic resistance,virulence,and adaptivemetabolicfunctions.[55][105]Although horizontal gene transfer is rare in eukaryotes, likely examples have been identified ofprotistandalgagenomes containing genes of bacterial origin.[106][107]
Genome
[edit]Thegenomeis the total genetic material of an organism and includes both the genes andnon-coding sequences.[108]Eukaryotic genes can be annotated using FINDER.[109]
Number of genes
[edit]
Thegenome size,and the number of genes it encodes varies widely between organisms. The smallest genomes occur inviruses,[118]andviroids(which act as a single non-coding RNA gene).[119]Conversely, plants can have extremely large genomes,[120]withricecontaining >46,000 protein-coding genes.[114]The total number of protein-coding genes (the Earth'sproteome) is estimated to be 5 million sequences.[121]
Although the number of base-pairs of DNA in the human genome has been known since the 1950s, the estimated number of genes has changed over time as definitions of genes, and methods of detecting them have been refined. Initial theoretical predictions of the number of human genes in the 1960s and 1970s were based on mutation load estimates and the numbers of mRNAs and these estimates tended to be about 30,000 protein-coding genes.[122][123][124]During the 1990s there were guesstimates of up to 100,000 genes and early data on detection of mRNAs (expressed sequence tags) suggested more than the traditional value of 30,000 genes that had been reported in the textbooks during the 1980s.[125]
The initial draft sequences of the human genome confirmed the earlier predictions of about 30,000 protein-coding genes however that estimate has fallen to about 19,000 with the ongoingGENCODEannotation project.[126]The number of noncoding genes is not known with certainty but the latest estimates from Ensembl suggest 26,000 noncoding genes.[127]
Essential genes
[edit]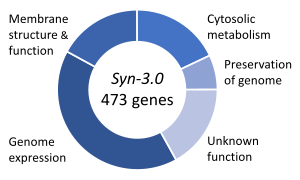
Essential genes are the set of genes thought to be critical for an organism's survival.[129]This definition assumes the abundant availability of all relevantnutrientsand the absence of environmental stress. Only a small portion of an organism's genes are essential. In bacteria, an estimated 250–400 genes are essential forEscherichia coliandBacillus subtilis,which is less than 10% of their genes.[130][131][132]Half of these genes areorthologsin both organisms and are largely involved inprotein synthesis.[132]In the budding yeastSaccharomyces cerevisiaethe number of essential genes is slightly higher, at 1000 genes (~20% of their genes).[133]Although the number is more difficult to measure in higher eukaryotes, mice and humans are estimated to have around 2000 essential genes (~10% of their genes).[134]The synthetic organism,Syn 3,has a minimal genome of 473 essential genes and quasi-essential genes (necessary for fast growth), although 149 have unknown function.[128]
Essential genes includehousekeeping genes(critical for basic cell functions)[135]as well as genes that are expressed at different times in the organismsdevelopmentorlife cycle.[136]Housekeeping genes are used asexperimental controlswhenanalysing gene expression,since they areconstitutively expressedat a relatively constant level.
Genetic and genomic nomenclature
[edit]Gene nomenclaturehas been established by theHUGO Gene Nomenclature Committee(HGNC), a committee of theHuman Genome Organisation,for each known human gene in the form of an approved gene name andsymbol(short-formabbreviation), which can be accessed through a database maintained by HGNC. Symbols are chosen to be unique, and each gene has only one symbol (although approved symbols sometimes change). Symbols are preferably kept consistent with other members of agene familyand with homologs in other species, particularly themousedue to its role as a commonmodel organism.[137]
Genetic engineering
[edit]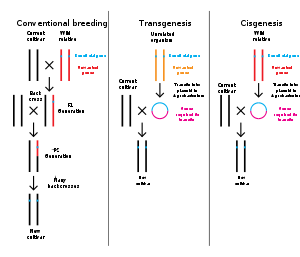
Genetic engineering is the modification of an organism'sgenomethroughbiotechnology.Since the 1970s, avariety of techniqueshave been developed to specifically add, remove and edit genes in an organism.[138]Recently developedgenome engineeringtechniques use engineerednucleaseenzymesto create targetedDNA repairin achromosometo either disrupt or edit a gene when the break is repaired.[139][140][141][142]The related termsynthetic biologyis sometimes used to refer to extensive genetic engineering of an organism.[143]
Genetic engineering is now a routine research tool withmodel organisms.For example, genes are easily added tobacteria[144]and lineages ofknockout micewith a specific gene's function disrupted are used to investigate that gene's function.[145][146]Many organisms have been genetically modified for applications inagriculture,industrial biotechnology, andmedicine.
For multicellular organisms, typically theembryois engineered which grows into the adultgenetically modified organism.[147]However, the genomes of cells in an adult organism can be edited usinggene therapytechniques to treat genetic diseases.
See also
[edit]References
[edit]Citations
[edit]- ^abcOrgogozo V, Peluffo AE, Morizot B (2016)."The" Mendelian Gene "and the" Molecular Gene ": Two Relevant Concepts of Genetic Units"(PDF).Current Topics in Developmental Biology.119:1–26.doi:10.1016/bs.ctdb.2016.03.002.PMID27282022.S2CID24583286.
- ^"What is a gene?: MedlinePlus Genetics".MedlinePlus.17 September 2020.Retrieved4 January2021.
- ^Hirsch ED (2002).The new dictionary of cultural literacy.Boston: Houghton Mifflin.ISBN0-618-22647-8.OCLC50166721.
- ^Elston RC, Satagopan JM, Sun S (2012). "Genetic terminology".Statistical Human Genetics.Methods in Molecular Biology. Vol. 850. Humana Press. pp. 1–9.doi:10.1007/978-1-61779-555-8_1.ISBN978-1-61779-554-1.PMC4450815.PMID22307690.
- ^abJohannsen W (1909).Elemente der exakten Erblichkeitslehre[Elements of the exact theory of heredity] (in German). Jena, Germany: Gustav Fischer. p. 124.From p. 124:"Dieses" etwas "in den Gameten bezw. in der Zygote,... – kurz, was wir eben Gene nennen wollen – bedingt sind."(This "something" in the gametes or in the zygote, which has crucial importance for the character of the organism, is usually called by the quite ambiguous termAnlagen[primordium, from the German wordAnlagefor "plan, arrangement; rough sketch" ]. Many other terms have been suggested, mostly unfortunately in closer connection with certain hypothetical opinions. The word "pangene", which was introduced by Darwin, is perhaps used most frequently in place ofAnlagen.However, the word "pangene" was not well chosen, as it is a compound word containing the rootspan(the neuter form of Πας all, every) andgen(from γί-γ(ε)ν-ομαι, to become). Only the meaning of this latter [i.e.,gen] comes into consideration here; just the basic idea – [namely,] that a trait in the developing organism can be determined or is influenced by "something" in the gametes – should find expression. No hypothesis about the nature of this "something" should be postulated or supported by it. For that reason it seems simplest to use in isolation the last syllablegenfrom Darwin's well-known word, which alone is of interest to us, in order to replace, with it, the poor, ambiguous wordAnlage.Thus we will say simply "gene" and "genes" for "pangene" and "pangenes". The word gene is completely free of any hypothesis; it expresses only the established fact that in any case many traits of the organism are determined by specific, separable, and thus independent "conditions", "foundations", "plans" – in short, precisely what we want to call genes.)
- ^abcdKampourakis K (2017).Making Sense of Genes.Cambridge, UK: Cambridge University Press.
- ^Gericke N, Hagberg M (5 December 2006). "Definition of historical models of gene function and their relation to students' understanding of genetics".Science & Education.16(7–8): 849–881.Bibcode:2007Sc&Ed..16..849G.doi:10.1007/s11191-006-9064-4.S2CID144613322.
- ^Meunier R (2022)."Stanford Encyclopedia of Philosophy: Gene".Stanford Encyclopedia of Philosophy.Retrieved28 February2023.
- ^Kellis M, Wold B, Snyder MP, Bernstein BE, Kundaje A, Marinov GK, et al. (April 2014)."Defining functional DNA elements in the human genome".Proceedings of the National Academy of Sciences of the United States of America.111(17): 6131–8.Bibcode:2014PNAS..111.6131K.doi:10.1073/pnas.1318948111.PMC4035993.PMID24753594.
- ^abDawkins R (1976).The selfish gene.Oxford, UK: Oxford University Press.
- ^Stoltz K, Griffiths P (2004)."Genes: Philosophical Analyses Put to the Test".History and Philosophy of the Life Sciences.26(1): 5–28.doi:10.1080/03919710412331341621.JSTOR23333378.PMID15791804.
- ^Beadle GW, Tatum EL (November 1941)."Genetic Control of Biochemical Reactions in Neurospora".Proceedings of the National Academy of Sciences of the United States of America.27(11): 499–506.Bibcode:1941PNAS...27..499B.doi:10.1073/pnas.27.11.499.PMC1078370.PMID16588492.
- ^Horowitz NH, Berg P, Singer M, Lederberg J, Susman M, Doebley J, Crow JF (January 2004)."A centennial: George W. Beadle, 1903-1989".Genetics.166(1): 1–10.doi:10.1534/genetics.166.1.1.PMC1470705.PMID15020400.
- ^Judson HF (1996).The Eight Day of Creation(Expanded ed.). Plainview, NY (US): Cold Spring Harbor Laboratory Press.
- ^abWatson JD (1965).Molecular Biology of the Gene.New York, NY, US: W.A. Benjamin, Inc.
- ^Alberts B, Bray D, Lewis J, Raff M, Roberts K, Watson JD (1994).Molecular Biology of the Cell: Third Edition.London, UK: Garland Publishing, Inc.ISBN0-8153-1619-4.
- ^Moran LA, Horton HR, Scrimgeour KG, Perry MD (2012).Principles of Biochemistry: Fifth Edition.Upper Saddle River, NJ, US: Pearson.
- ^Lewin B (2004).Genes VIII.Upper Saddle River, NJ, US: Pearson/Prentice Hall.
- ^Piovesan A, Pelleri MC, Antonaros F, Strippoli P, Caracausi M, and Vitale L (2019)."On the length, weight and GC content of the human genome".BMC Research Notes.12(1): 106–173.doi:10.1186/s13104-019-4137-z.PMC6391780.PMID30813969.
- ^Hubé F, and Francastel C (2015)."Mammalian Introns: When the Junk Generates Molecular Diversity".International Journal of Molecular Sciences.16(3): 4429–4452.doi:10.3390/ijms16034429.PMC4394429.PMID25710723.
- ^Francis WR, and Wörheide G (2017)."Similar ratios of introns to intergenic sequence across animal genomes".Genome Biology and Evolution.9(6): 1582–1598.doi:10.1093/gbe/evx103.PMC5534336.PMID28633296.
- ^Mortola E, Long M (2021)."Turning Junk into Us: How Genes Are Born".American Scientist.109:174–182.
- ^Hopkin K (2009). "The Evolving Definition of a Gene: With the discovery that nearly all of the genome is transcribed, the definition of a" gene "needs another revision".BioScience.59:928–931.doi:10.1525/bio.2009.59.11.3.S2CID88157272.
- ^Pearson H (2006)."What Is a Gene?".Nature.441(7092): 399–401.Bibcode:2006Natur.441..398P.doi:10.1038/441398a.PMID16724031.S2CID4420674.
- ^Pennisi E (2007)."DNA study forces rethink of what it means to be a gene".Science.316(5831): 1556–1557.doi:10.1126/science.316.5831.1556.PMID17569836.S2CID36463252.
- ^Wolf YI, Kazlauskas D, Iranzo J, Lucía-Sanz A, Kuhn JH, Krupovic M, et al. (November 2018). Racaniello VR (ed.)."Origins and Evolution of the Global RNA Virome".mBio.9(6). Eric Delwart, Luis Enjuanes: e02329–18.doi:10.1128/mBio.02329-18.PMC6282212.PMID30482837.
- ^abPennisi E(June 2007)."Genomics. DNA study forces rethink of what it means to be a gene".Science.316(5831): 1556–7.doi:10.1126/science.316.5831.1556.PMID17569836.S2CID36463252.
- ^Marande W, Burger G (October 2007). "Mitochondrial DNA as a genomic jigsaw puzzle".Science.318(5849). AAAS: 415.Bibcode:2007Sci...318..415M.doi:10.1126/science.1148033.PMID17947575.S2CID30948765.
- ^Parra G, Reymond A, Dabbouseh N, Dermitzakis ET, Castelo R, Thomson TM, et al. (January 2006)."Tandem chimerism as a means to increase protein complexity in the human genome".Genome Research.16(1): 37–44.doi:10.1101/gr.4145906.PMC1356127.PMID16344564.
- ^abcGerstein MB, Bruce C, Rozowsky JS, Zheng D, Du J, Korbel JO, et al. (June 2007)."What is a gene, post-ENCODE? History and updated definition".Genome Research.17(6): 669–81.doi:10.1101/gr.6339607.PMID17567988.
- ^Noble D (September 2008)."Genes and causation".Philosophical Transactions. Series A, Mathematical, Physical, and Engineering Sciences.366(1878): 3001–15.Bibcode:2008RSPTA.366.3001N.doi:10.1098/rsta.2008.0086.PMID18559318.
- ^"Blending Inheritance - an overview | ScienceDirect Topics".
- ^"genesis".Oxford English Dictionary(Online ed.).Oxford University Press.(Subscription orparticipating institution membershiprequired.)
- ^Magner LN (2002).A History of the Life Sciences(Third ed.).Marcel Dekker,CRC Press.p. 371.ISBN978-0-203-91100-6.
- ^Henig RM (2000).The Monk in the Garden: The Lost and Found Genius of Gregor Mendel, the Father of Genetics.Boston: Houghton Mifflin. pp.1–9.ISBN978-0395-97765-1.
- ^abde Vries H(1889).Intracellulare Pangenese[Intracellular Pangenesis] (in German). Translated byGager CS.Jena: Verlag von Gustav Fischer.Translated in 1908 from German to English by Open Court Publishing Co., Chicago, 1910
- ^Bateson W (1906)."The progress of genetic research".In Wilks W (ed.).Report of the Third International Conference 1906 on Genetics.London, England: Royal Horticultural Society. pp. 90–97.
... the science itself [i.e. the study of the breeding and hybridisation of plants] is still nameless, and we can only describe our pursuit by cumbrous and often misleading periphrasis. To meet this difficulty I suggest for the consideration of this Congress the termGenetics,which sufficiently indicates that our labors are devoted to the elucidation of the phenomena of heredity and variation: in other words, to the physiology of Descent, with implied bearing on the theoretical problems of the evolutionist and the systematist, and application to the practical problems of breeders, whether of animals or plants.
- ^Avery OT, Macleod CM, McCarty M (February 1944)."Studies on the Chemical Nature of the Substance Inducing Transformation of Pneumococcal Types: Induction of Transformation by a Desoxyribonucleic Acid Fraction Isolated From Pneumococcus Type III".The Journal of Experimental Medicine.79(2): 137–58.doi:10.1084/jem.79.2.137.PMC2135445.PMID19871359.Reprint:Avery OT, MacLeod CM, McCarty M (February 1979)."Studies on the chemical nature of the substance inducing transformation of pneumococcal types. Inductions of transformation by a desoxyribonucleic acid fraction isolated from pneumococcus type III".The Journal of Experimental Medicine.149(2): 297–326.doi:10.1084/jem.149.2.297.PMC2184805.PMID33226.
- ^Hershey AD, Chase M (May 1952)."Independent functions of viral protein and nucleic acid in growth of bacteriophage".The Journal of General Physiology.36(1): 39–56.doi:10.1085/jgp.36.1.39.PMC2147348.PMID12981234.
- ^Judson H(1979).The Eighth Day of Creation: Makers of the Revolution in Biology.Cold Spring Harbor Laboratory Press. pp. 51–169.ISBN978-0-87969-477-7.
- ^Watson JD, Crick FH (April 1953)."Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid"(PDF).Nature.171(4356): 737–8.Bibcode:1953Natur.171..737W.doi:10.1038/171737a0.PMID13054692.S2CID4253007.
- ^Benzer S (June 1955)."Fine Structure of a Genetic Region in Bacteriophage".Proceedings of the National Academy of Sciences of the United States of America.41(6): 344–54.Bibcode:1955PNAS...41..344B.doi:10.1073/pnas.41.6.344.PMC528093.PMID16589677.
- ^Benzer S (November 1959)."On the Topology of the Genetic Fine Structure".Proceedings of the National Academy of Sciences of the United States of America.45(11): 1607–20.Bibcode:1959PNAS...45.1607B.doi:10.1073/pnas.45.11.1607.PMC222769.PMID16590553.
- ^Min Jou W, Haegeman G, Ysebaert M, Fiers W (May 1972). "Nucleotide sequence of the gene coding for the bacteriophage MS2 coat protein".Nature.237(5350): 82–8.Bibcode:1972Natur.237...82J.doi:10.1038/237082a0.PMID4555447.S2CID4153893.
- ^Sanger F, Nicklen S, Coulson AR (December 1977)."DNA sequencing with chain-terminating inhibitors".Proceedings of the National Academy of Sciences of the United States of America.74(12): 5463–7.Bibcode:1977PNAS...74.5463S.doi:10.1073/pnas.74.12.5463.PMC431765.PMID271968.
- ^Adams JU (2008)."DNA Sequencing Technologies".Nature Education Knowledge.SciTable.1(1). Nature Publishing Group: 193.
- ^Huxley J (1942).Evolution: the Modern Synthesis.Cambridge, Massachusetts: MIT Press.ISBN978-0262513661.
- ^Williams GC (2001).Adaptation and Natural Selection a Critique of Some Current Evolutionary Thought(Online ed.). Princeton: Princeton University Press.ISBN9781400820108.
- ^Dawkins R (1989).The extended phenotype(Paperback ed.). Oxford: Oxford University Press.ISBN978-0-19-286088-0.
- ^Duret L (2008)."Neutral Theory: The Null Hypothesis of Molecular Evolution".Nature Education.1:218.
- ^abcdefghijklmnopqrstuvwxyzaaabacadaeafagahaiAlberts B,Johnson A,Lewis J,Raff M,Roberts K,Walter P(2002).Molecular Biology of the Cell(Fourth ed.). New York: Garland Science.ISBN978-0-8153-3218-3.
- ^Stryer L, Berg JM, Tymoczko JL (2002).Biochemistry(5th ed.). San Francisco: W.H. Freeman.ISBN978-0-7167-4955-4.
- ^Bolzer A, Kreth G, Solovei I, Koehler D, Saracoglu K, Fauth C, et al. (May 2005)."Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes".PLOS Biology.3(5): e157.doi:10.1371/journal.pbio.0030157.PMC1084335.PMID15839726.

- ^Braig M, Schmitt CA (March 2006)."Oncogene-induced senescence: putting the brakes on tumor development".Cancer Research.66(6): 2881–4.doi:10.1158/0008-5472.CAN-05-4006.PMID16540631.
- ^abBennett PM (March 2008)."Plasmid encoded antibiotic resistance: acquisition and transfer of antibiotic resistance genes in bacteria".British Journal of Pharmacology.153(Suppl 1): S347-57.doi:10.1038/sj.bjp.0707607.PMC2268074.PMID18193080.
- ^International Human Genome Sequencing Consortium (October 2004)."Finishing the euchromatic sequence of the human genome".Nature.431(7011): 931–45.Bibcode:2004Natur.431..931H.doi:10.1038/nature03001.PMID15496913.
- ^abShafee, Thomas; Lowe, Rohan (2017)."Eukaryotic and prokaryotic gene structure".WikiJournal of Medicine.4(1).doi:10.15347/wjm/2017.002.ISSN2002-4436.
- ^Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (July 2008). "Mapping and quantifying mammalian transcriptomes by RNA-Seq".Nature Methods.5(7): 621–8.doi:10.1038/nmeth.1226.PMID18516045.S2CID205418589.
- ^Pennacchio LA, Bickmore W, Dean A, Nobrega MA, Bejerano G (April 2013)."Enhancers: five essential questions".Nature Reviews. Genetics.14(4): 288–95.doi:10.1038/nrg3458.PMC4445073.PMID23503198.
- ^Maston GA, Evans SK, Green MR (2006)."Transcriptional regulatory elements in the human genome".Annual Review of Genomics and Human Genetics.7:29–59.doi:10.1146/annurev.genom.7.080505.115623.PMID16719718.
- ^Mignone F, Gissi C, Liuni S, Pesole G (28 February 2002)."Untranslated regions of mRNAs".Genome Biology.3(3): REVIEWS0004.doi:10.1186/gb-2002-3-3-reviews0004.PMC139023.PMID11897027.
- ^Bicknell AA, Cenik C, Chua HN, Roth FP, Moore MJ (December 2012)."Introns in UTRs: why we should stop ignoring them".BioEssays.34(12): 1025–34.doi:10.1002/bies.201200073.PMID23108796.S2CID5808466.
- ^Salgado H, Moreno-Hagelsieb G, Smith TF, Collado-Vides J (June 2000)."Operons in Escherichia coli: genomic analyses and predictions".Proceedings of the National Academy of Sciences of the United States of America.97(12): 6652–7.Bibcode:2000PNAS...97.6652S.doi:10.1073/pnas.110147297.PMC18690.PMID10823905.
- ^Blumenthal T (November 2004)."Operons in eukaryotes".Briefings in Functional Genomics & Proteomics.3(3): 199–211.doi:10.1093/bfgp/3.3.199.PMID15642184.
- ^Jacob F, Monod J (June 1961). "Genetic regulatory mechanisms in the synthesis of proteins".Journal of Molecular Biology.3(3): 318–56.doi:10.1016/S0022-2836(61)80072-7.PMID13718526.S2CID19804795.
- ^Pozzoli U, Menozzi G, Comi GP, Cagliani R, Bresolin N, Sironi M (January 2007). "Intron size in mammals: complexity comes to terms with economy".Trends in Genetics.23(1): 20–24.doi:10.1016/j.tig.2006.10.003.PMID17070957.
- ^Marais G, Nouvellet P, Keightley PD, Charlesworth B (May 2005)."Intron size and exon evolution in Drosophila".Genetics.170(1): 481–485.doi:10.1534/genetics.104.037333.PMC1449718.PMID15781704.
- ^Kumar A (September 2009)."An overview of nested genes in eukaryotic genomes".Eukaryotic Cell.8(9): 1321–1329.doi:10.1128/EC.00143-09.PMC2747821.PMID19542305..
- ^Spilianakis CG, Lalioti MD, Town T, Lee GR, Flavell RA (June 2005). "Interchromosomal associations between alternatively expressed loci".Nature.435(7042): 637–645.Bibcode:2005Natur.435..637S.doi:10.1038/nature03574.PMID15880101.S2CID1755326.
- ^Williams A, Spilianakis CG, Flavell RA (April 2010)."Interchromosomal association and gene regulation in trans".Trends in Genetics.26(4): 188–197.doi:10.1016/j.tig.2010.01.007.PMC2865229.PMID20236724.
- ^Lei Q, Li C, Zuo Z, Huang C, Cheng H, Zhou R (March 2016)."Evolutionary Insights into RNA trans-Splicing in Vertebrates".Genome Biology and Evolution.8(3): 562–577.doi:10.1093/gbe/evw025.PMC4824033.PMID26966239.
- ^Wright BW, Molloy MP, Jaschke PR (March 2022)."Overlapping genes in natural and engineered genomes".Nature Reviews. Genetics.23(3): 154–168.doi:10.1038/s41576-021-00417-w.PMC8490965.PMID34611352.
- ^abEddy SR (December 2001). "Non-coding RNA genes and the modern RNA world".Nature Reviews. Genetics.2(12): 919–29.doi:10.1038/35103511.PMID11733745.S2CID18347629.
- ^Crick FH, Barnett L, Brenner S, Watts-Tobin RJ (December 1961). "General nature of the genetic code for proteins".Nature.192(4809): 1227–32.Bibcode:1961Natur.192.1227C.doi:10.1038/1921227a0.PMID13882203.S2CID4276146.
- ^Crick FH (October 1962)."The genetic code".Scientific American.207(4). WH Freeman and Company: 66–74.Bibcode:1962SciAm.207d..66C.doi:10.1038/scientificamerican1062-66.PMID13882204.
- ^Woodson SA (May 1998)."Ironing out the kinks: splicing and translation in bacteria".Genes & Development.12(9): 1243–7.doi:10.1101/gad.12.9.1243.PMID9573040.
- ^Jacob F,Monod J(June 1961). "Genetic regulatory mechanisms in the synthesis of proteins".Journal of Molecular Biology.3(3): 318–56.doi:10.1016/S0022-2836(61)80072-7.PMID13718526.S2CID19804795.
- ^Koonin EV, Dolja VV (January 1993). "Evolution and taxonomy of positive-strand RNA viruses: implications of comparative analysis of amino acid sequences".Critical Reviews in Biochemistry and Molecular Biology.28(5): 375–430.doi:10.3109/10409239309078440.PMID8269709.
- ^Domingo E (2001). "RNA Virus Genomes".eLS.doi:10.1002/9780470015902.a0001488.pub2.ISBN978-0470016176.
- ^Domingo E, Escarmís C, Sevilla N, Moya A, Elena SF, Quer J, et al. (June 1996)."Basic concepts in RNA virus evolution".FASEB Journal.10(8): 859–64.doi:10.1096/fasebj.10.8.8666162.PMID8666162.S2CID20865732.
- ^Miko I (2008)."Gregor Mendel and the Principles of Inheritance".Nature Education Knowledge.SciTable.1(1). Nature Publishing Group: 134.
- ^Chial H (2008)."Mendelian Genetics: Patterns of Inheritance and Single-Gene Disorders".Nature Education Knowledge.SciTable.1(1). Nature Publishing Group: 63.
- ^McCarthy D, Minner C, Bernstein H, Bernstein C (October 1976). "DNA elongation rates and growing point distributions of wild-type phage T4 and a DNA-delay amber mutant".Journal of Molecular Biology.106(4): 963–81.doi:10.1016/0022-2836(76)90346-6.PMID789903.
- ^abLobo I, Shaw K (2008)."Discovery and Types of Genetic Linkage".Nature Education Knowledge.SciTable.1(1). Nature Publishing Group: 139.
- ^Nachman MW, Crowell SL (September 2000)."Estimate of the Mutation Rate per Nucleotide in Humans".Genetics.156(1): 297–304.doi:10.1093/genetics/156.1.297.PMC1461236.PMID10978293.
- ^Roach JC, Glusman G, Smit AF, Huff CD, Hubley R, Shannon PT, et al. (April 2010)."Analysis of genetic inheritance in a family quartet by whole-genome sequencing".Science.328(5978): 636–9.Bibcode:2010Sci...328..636R.doi:10.1126/science.1186802.PMC3037280.PMID20220176.
- ^Drake JW, Charlesworth B, Charlesworth D, Crow JF (April 1998)."Rates of Spontaneous Mutation".Genetics.148(4): 1667–86.doi:10.1093/genetics/148.4.1667.PMC1460098.PMID9560386.
- ^Pyeritz, Reed E., Bruce R. Korf, and Wayne W. Grody, eds. Emery and Rimoin’s principles and practice of medical genetics and genomics: foundations. Academic Press, 2018.
- ^"What kinds of gene mutations are possible?".Genetics Home Reference.United States National Library of Medicine. 11 May 2015.Retrieved19 May2015.
- ^Andrews CA (2010)."Natural Selection, Genetic Drift, and Gene Flow Do Not Act in Isolation in Natural Populations".Nature Education Knowledge.SciTable.3(10). Nature Publishing Group: 5.
- ^Patterson C (November 1988)."Homology in classical and molecular biology".Molecular Biology and Evolution.5(6): 603–25.doi:10.1093/oxfordjournals.molbev.a040523.PMID3065587.
- ^Graur D (2016).Molecular and Genome Evolution.Sunderland MA (US): Sinauer Associates, Inc.ISBN9781605354699.
- ^Graur D (2016).Molecular and Genome Evolution.Sunderland MA (US): Sinauer Associates, Inc.ISBN9781605354699.
- ^Jensen RA (2001)."Orthologs and paralogs - we need to get it right".Genome Biology.2(8): INTERACTIONS1002.doi:10.1186/gb-2001-2-8-interactions1002.PMC138949.PMID11532207.
- ^Studer RA, Robinson-Rechavi M (May 2009)."How confident can we be that orthologs are similar, but paralogs differ?".Trends in Genetics.25(5): 210–6.doi:10.1016/j.tig.2009.03.004.PMID19368988.
- ^Altenhoff AM, Studer RA, Robinson-Rechavi M, Dessimoz C (2012)."Resolving the ortholog conjecture: orthologs tend to be weakly, but significantly, more similar in function than paralogs".PLOS Computational Biology.8(5): e1002514.Bibcode:2012PLSCB...8E2514A.doi:10.1371/journal.pcbi.1002514.PMC3355068.PMID22615551.

- ^abGuerzoni D, McLysaght A (November 2011)."De novo origins of human genes".PLOS Genetics.7(11): e1002381.doi:10.1371/journal.pgen.1002381.PMC3213182.PMID22102832.

- ^Reams AB, Roth JR (February 2015)."Mechanisms of gene duplication and amplification".Cold Spring Harbor Perspectives in Biology.7(2): a016592.doi:10.1101/cshperspect.a016592.PMC4315931.PMID25646380.
- ^Demuth JP, De Bie T, Stajich JE, Cristianini N, Hahn MW (December 2006)."The evolution of mammalian gene families".PLOS ONE.1(1): e85.Bibcode:2006PLoSO...1...85D.doi:10.1371/journal.pone.0000085.PMC1762380.PMID17183716.

- ^Knowles DG, McLysaght A (October 2009)."Recent de novo origin of human protein-coding genes".Genome Research.19(10): 1752–9.doi:10.1101/gr.095026.109.PMC2765279.PMID19726446.
- ^Wu DD, Irwin DM, Zhang YP (November 2011)."De novo origin of human protein-coding genes".PLOS Genetics.7(11): e1002379.doi:10.1371/journal.pgen.1002379.PMC3213175.PMID22102831.

- ^McLysaght A, Guerzoni D (September 2015)."New genes from non-coding sequence: the role of de novo protein-coding genes in eukaryotic evolutionary innovation".Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences.370(1678): 20140332.doi:10.1098/rstb.2014.0332.PMC4571571.PMID26323763.
- ^Neme R, Tautz D (February 2013)."Phylogenetic patterns of emergence of new genes support a model of frequent de novo evolution".BMC Genomics.14(1): 117.doi:10.1186/1471-2164-14-117.PMC3616865.PMID23433480.
- ^Treangen TJ, Rocha EP (January 2011)."Horizontal transfer, not duplication, drives the expansion of protein families in prokaryotes".PLOS Genetics.7(1): e1001284.doi:10.1371/journal.pgen.1001284.PMC3029252.PMID21298028.

- ^Ochman H, Lawrence JG, Groisman EA (May 2000). "Lateral gene transfer and the nature of bacterial innovation".Nature.405(6784): 299–304.Bibcode:2000Natur.405..299O.doi:10.1038/35012500.PMID10830951.S2CID85739173.
- ^Keeling PJ, Palmer JD (August 2008). "Horizontal gene transfer in eukaryotic evolution".Nature Reviews. Genetics.9(8): 605–18.doi:10.1038/nrg2386.PMID18591983.S2CID213613.
- ^Schönknecht G, Chen WH, Ternes CM, Barbier GG, Shrestha RP, Stanke M, et al. (March 2013)."Gene transfer from bacteria and archaea facilitated evolution of an extremophilic eukaryote".Science.339(6124): 1207–10.Bibcode:2013Sci...339.1207S.doi:10.1126/science.1231707.PMID23471408.S2CID5502148.
- ^Ridley, M. (2006).Genome.New York, NY: Harper Perennial.ISBN0-06-019497-9
- ^Banerjee S, Bhandary P, Woodhouse M, Sen TZ, Wise RP, Andorf CM (April 2021)."FINDER: an automated software package to annotate eukaryotic genes from RNA-Seq data and associated protein sequences".BMC Bioinformatics.44(9): e89.doi:10.1186/s12859-021-04120-9.PMC8056616.PMID33879057.
- ^Watson, JD, Baker TA, Bell SP, Gann A, Levine M, Losick R. (2004). "Ch9-10", Molecular Biology of the Gene, 5th ed., Peason Benjamin Cummings; CSHL Press.
- ^"Integr8 – A.thaliana Genome Statistics".
- ^"Understanding the Basics".The Human Genome Project.Retrieved26 April2015.
- ^"WS227 Release Letter".WormBase. 10 August 2011. Archived fromthe originalon 28 November 2013.Retrieved19 November2013.
- ^abYu J, Hu S, Wang J, Wong GK, Li S, Liu B, et al. (April 2002). "A draft sequence of the rice genome (Oryza sativa L. ssp. indica)".Science.296(5565): 79–92.Bibcode:2002Sci...296...79Y.doi:10.1126/science.1068037.PMID11935017.S2CID208529258.
- ^Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J, et al. (April 1981). "Sequence and organization of the human mitochondrial genome".Nature.290(5806): 457–65.Bibcode:1981Natur.290..457A.doi:10.1038/290457a0.PMID7219534.S2CID4355527.
- ^Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG, et al. (March 2000). "The genome sequence of Drosophila melanogaster".Science.287(5461): 2185–95.Bibcode:2000Sci...287.2185..CiteSeerX10.1.1.549.8639.doi:10.1126/science.287.5461.2185.PMID10731132.
- ^Pertea M, Salzberg SL (2010)."Between a chicken and a grape: estimating the number of human genes".Genome Biology.11(5): 206.doi:10.1186/gb-2010-11-5-206.PMC2898077.PMID20441615.
- ^Belyi VA, Levine AJ, Skalka AM (December 2010)."Sequences from ancestral single-stranded DNA viruses in vertebrate genomes: the parvoviridae and circoviridae are more than 40 to 50 million years old".Journal of Virology.84(23): 12458–62.doi:10.1128/JVI.01789-10.PMC2976387.PMID20861255.
- ^Flores R, Di Serio F, Hernández C (February 1997). "Viroids: The Noncoding Genomes".Seminars in Virology.8(1): 65–73.doi:10.1006/smvy.1997.0107.
- ^Zonneveld BJ (2010)."New Record Holders for Maximum Genome Size in Eudicots and Monocots".Journal of Botany.2010:1–4.doi:10.1155/2010/527357.
- ^Perez-Iratxeta C, Palidwor G, Andrade-Navarro MA (December 2007)."Towards completion of the Earth's proteome".EMBO Reports.8(12): 1135–41.doi:10.1038/sj.embor.7401117.PMC2267224.PMID18059312.
- ^Muller HJ (1966)."The gene material as the initiator and the organizing basis of life".American Naturalist.100(915): 493–517.doi:10.1086/282445.JSTOR2459205.S2CID84202145.
- ^Ohno S (1972). "So much" junk "DNA in our genome".Brookhaven Symposia in Biology.23:366–370.PMID5065367.
- ^Hatje K, Mühlhausen S, Simm D, Killmar M (2019)."The Protein-Coding Human Genome: Annotating High-Hanging Fruits".BioEssays.41(11): 1900066.doi:10.1002/bies.201900066.PMID31544971.S2CID202732556.
- ^Schuler GD,Boguski MS,Stewart EA, Stein LD, Gyapay G, Rice K, et al. (October 1996). "A gene map of the human genome".Science.274(5287): 540–6.Bibcode:1996Sci...274..540S.doi:10.1126/science.274.5287.540.PMID8849440.S2CID22619.
- ^Chi KR (October 2016)."The dark side of the human genome".Nature.538(7624): 275–277.Bibcode:2016Natur.538..275C.doi:10.1038/538275a.PMID27734873.
- ^"Human assembly and gene annotation".Ensembl.2022.Retrieved28 February2023.
- ^abHutchison CA, Chuang RY, Noskov VN, Assad-Garcia N, Deerinck TJ, Ellisman MH, et al. (March 2016)."Design and synthesis of a minimal bacterial genome".Science.351(6280): aad6253.Bibcode:2016Sci...351.....H.doi:10.1126/science.aad6253.PMID27013737.
- ^Glass JI, Assad-Garcia N, Alperovich N, Yooseph S, Lewis MR, Maruf M, et al. (January 2006)."Essential genes of a minimal bacterium".Proceedings of the National Academy of Sciences of the United States of America.103(2): 425–30.Bibcode:2006PNAS..103..425G.doi:10.1073/pnas.0510013103.PMC1324956.PMID16407165.
- ^Gerdes SY, Scholle MD, Campbell JW, Balázsi G, Ravasz E, Daugherty MD, et al. (October 2003)."Experimental determination and system level analysis of essential genes in Escherichia coli MG1655".Journal of Bacteriology.185(19): 5673–84.doi:10.1128/jb.185.19.5673-5684.2003.PMC193955.PMID13129938.
- ^Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, et al. (2006)."Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection".Molecular Systems Biology.2:2006.0008.doi:10.1038/msb4100050.PMC1681482.PMID16738554.
- ^abJuhas M, Reuß DR, Zhu B, Commichau FM (November 2014)."Bacillus subtilis and Escherichia coli essential genes and minimal cell factories after one decade of genome engineering".Microbiology.160(Pt 11): 2341–2351.doi:10.1099/mic.0.079376-0.PMID25092907.
- ^Tu Z, Wang L, Xu M, Zhou X, Chen T, Sun F (February 2006)."Further understanding human disease genes by comparing with housekeeping genes and other genes".BMC Genomics.7:31.doi:10.1186/1471-2164-7-31.PMC1397819.PMID16504025.

- ^Georgi B, Voight BF, Bućan M (May 2013)."From mouse to human: evolutionary genomics analysis of human orthologs of essential genes".PLOS Genetics.9(5): e1003484.doi:10.1371/journal.pgen.1003484.PMC3649967.PMID23675308.

- ^Eisenberg E, Levanon EY (October 2013). "Human housekeeping genes, revisited".Trends in Genetics.29(10): 569–74.doi:10.1016/j.tig.2013.05.010.PMID23810203.
- ^Amsterdam A, Hopkins N (September 2006). "Mutagenesis strategies in zebrafish for identifying genes involved in development and disease".Trends in Genetics.22(9): 473–8.doi:10.1016/j.tig.2006.06.011.PMID16844256.
- ^"About the HGNC".HGNC Database of Human Gene Names.HUGO Gene Nomenclature Committee. Archived fromthe originalon 26 March 2023.Retrieved14 May2015.
- ^Cohen SN, Chang AC (May 1973)."Recircularization and autonomous replication of a sheared R-factor DNA segment in Escherichia coli transformants".Proceedings of the National Academy of Sciences of the United States of America.70(5): 1293–7.Bibcode:1973PNAS...70.1293C.doi:10.1073/pnas.70.5.1293.PMC433482.PMID4576014.
- ^Esvelt KM, Wang HH (2013)."Genome-scale engineering for systems and synthetic biology".Molecular Systems Biology.9(1): 641.doi:10.1038/msb.2012.66.PMC3564264.PMID23340847.
- ^Tan WS, Carlson DF, Walton MW, Fahrenkrug SC, Hackett PB (2012). "Precision editing of large animal genomes".Advances in Genetics Volume 80.Vol. 80. pp. 37–97.doi:10.1016/B978-0-12-404742-6.00002-8.ISBN9780124047426.PMC3683964.PMID23084873.
- ^Puchta H, Fauser F (2013)."Gene targeting in plants: 25 years later".The International Journal of Developmental Biology.57(6–8): 629–37.doi:10.1387/ijdb.130194hp.PMID24166445.
- ^Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (November 2013)."Genome engineering using the CRISPR-Cas9 system".Nature Protocols.8(11): 2281–2308.doi:10.1038/nprot.2013.143.PMC3969860.PMID24157548.
- ^Kittleson JT, Wu GC, Anderson JC (August 2012). "Successes and failures in modular genetic engineering".Current Opinion in Chemical Biology.16(3–4): 329–36.doi:10.1016/j.cbpa.2012.06.009.PMID22818777.
- ^Berg P, Mertz JE (January 2010)."Personal reflections on the origins and emergence of recombinant DNA technology".Genetics.184(1): 9–17.doi:10.1534/genetics.109.112144.PMC2815933.PMID20061565.
- ^Austin CP, Battey JF, Bradley A, Bucan M, Capecchi M, Collins FS, et al. (September 2004)."The knockout mouse project".Nature Genetics.36(9): 921–4.doi:10.1038/ng0904-921.PMC2716027.PMID15340423.
- ^Guan C, Ye C, Yang X, Gao J (February 2010). "A review of current large-scale mouse knockout efforts".Genesis.48(2): 73–85.doi:10.1002/dvg.20594.PMID20095055.S2CID34470273.
- ^Deng C (October 2007)."In celebration of Dr. Mario R. Capecchi's Nobel Prize".International Journal of Biological Sciences.3(7): 417–9.doi:10.7150/ijbs.3.417.PMC2043165.PMID17998949.
Sources
[edit]- Main textbook
- Alberts B,Johnson A,Lewis J,Raff M,Roberts K,Walter P(2002).Molecular Biology of the Cell(Fourth ed.). New York: Garland Science.ISBN978-0-8153-3218-3.– A molecular biology textbook available free online through NCBI Bookshelf.

Further reading
[edit]- Watson JD,Baker TA,Bell SP, Gann A,Levine M,Losick R (2013).Molecular Biology of the Gene(7th ed.). Benjamin Cummings.ISBN978-0-321-90537-6.
- Dawkins R(1990).The Selfish Gene.Oxford University Press.ISBN978-0-19-286092-7.
- Ridley M(1999).Genome: The Autobiography of a Species in 23 Chapters.Fourth Estate.ISBN978-0-00-763573-3.
- Brown T (2002).Genomes(2nd ed.). New York: Wiley-Liss.ISBN978-0-471-25046-3.PMID20821850.
External links
[edit]- Comparative Toxicogenomics Database
- DNA From The Beginning – a primer on genes and DNA
- Gene – a searchable database of genes
- Genes– an Open Access journal
- IDconverter – converts gene IDs between public databases
- iHOP – Information Hyperlinked over Proteins
- TranscriptomeBrowser – Gene expression profile analysis
- The Protein Naming Utility, a database to identify and correct deficient gene names
- IMPC (International Mouse Phenotyping Consortium)– Encyclopedia of mammalian gene function
- Global Genes Project– Leading non-profit organization supporting people living with genetic diseases
- Encode threads explorer,Nature